SAMSON has a powerful Node Specification Language (NSL) that may be used to select data graph nodes based on their properties. For example, the Find command of the user interface of SAMSON lets users enter a NSL string to select nodes from the document:

A NSL expression may also be used to filter nodes from the document view. In the example below, the user has entered n.t sc (node.type sidechain) and hit the Enter key to select all side chains:
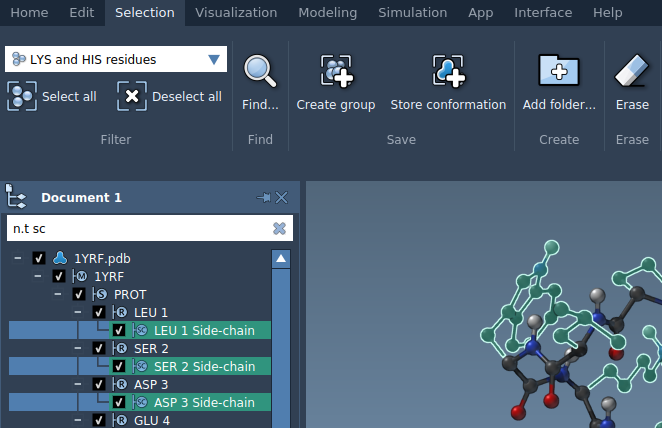
Here are some examples of NSL expressions:
Hydrogen: select all hydrogens (short version:H)atom.chainID > 2: select all atoms with a chain ID strictly larger than 2 (short version:a.ci > 2)Carbon in node.selected: select all carbons in the current selection (short version:C in n.s)bond.order > 1.5: select all bonds with order strictly larger than 1.5 (short version:b.o > 1.5)node.type backbone: select all backbone nodes (short version:n.t bb)O in node.type sidechain: select all oxygens in sidechain nodes (short version:O in n.t sc)"CA" within 5A of S: select all nodes named "CA" that are within 5 angstrom of any sulfur atom (short version:"CA" w 5A of S) (use quotes, since names may contain spaces)node.type residue beyond 5A of node.selected: select all residue nodes beyond 5 angstrom of the current selection (short version:n.t r b 5A of n.s)residue.secondaryStructure helix: select residue nodes in alpha helices (short version:r.ss h)node.type sidechain having S: select sidechain nodes that have at least one sulfur atom (short version:n.t sc h S)H linking O: select all hydrogens bonded to oxygen atoms (short version:H l O)
Logical operators may be used:
C or H: select atoms that are carbons or hydrogens
When using the Find command, selections may be saved as groups in the data graph:
"Interface" = ((n.t a in "A") w 4A of "B") or ((n.t a in "B") w 4A of "A"): creates a group containing atoms at the interface between"A"and"B"
When using the Find command as well, you may press the Tab key in the search box for context-sensitive completion. This is particularly useful when searching nodes by name. For example, entering "ALA (with the quote sign, to indicate a name) and pressing the Tab key will list all nodes whose name begins with "ALA, e.g:
"ALA 22 Backbone""ALA 22 Side-chain""ALA 22""ALA 28 Backbone""ALA 28 Side-chain""ALA 28"...
Specification by attributes
Nodes may be specified by their attributes. Since nodes may have different types in SAMSON (e.g. atom, bond, etc.), each attribute is defined in an attribute space.
The node attribute space (short name n) corresponds to attributes that are defined for each node. For example, the selection flag is a node attribute, since each node has a selection flag. Hence, the NSL expression node.selectionFlag true may match any node whose selection flag is true, regardless of its node type (atom, bond, etc.).
SAMSON's Node Specification Language supports four different attribute spaces: node (short name n), atom (short name a), bond (short name b), and residue (short name r).
Node attributes
Node attributes, defined in the node attribute space (short name n), correspond to attributes defined in each node of the data graph. For example, the selection flag is a node attribute, since each node has a selection flag. Hence, the NSL expression node.selectionFlag true may match any node whose selection flag is true, regardless of its node type (atom, bond, etc.).
Possible node attributes are:
hidden(short nameh)selected(short names)selectionFlag(short namesf)type(short namet)visibilityFlag(short namevf)visible(short namev)
hidden
The hidden attribute (short name h) matches nodes that are hidden, either because their visibility flag is false, or because the visibility flag of one of their ancestors is false.
Possible values: none.
Example:
n.h: matches all hidden nodes
selected
The selected attribute (short name s) matches nodes that are selected, either because their selection flag is true, or because the selection flag of one of their ancestors is true.
Possible values: none.
Example:
n.s: matches all selected nodes
selectionFlag
The selectionFlag attribute (short name sf) matches nodes based on their selection flag.
Possible values: true or false.
Example:
n.sf true: matches all nodes with a selection flag set totrue
type
The type attribute (short name t) matches nodes by type.
Possible values:
atom(short namea)camera(short nameca)backbone(short namebb)bond(short nameb)chain(short namec)conformation(short nameco)document(short named)dynamicalModelParticleSystem(short namedmps)interactionModelParticleSystem(short nameimps)label(short namela)folder(short namef)molecule(short namem)nodeGroup(short nameng)propertyModel(short namepm)pseudoatom(short namepa)residue(short namer)segment(short names)sidechain(short namesc)simulatorParticleSystem(short namesps)stateUpdaterParticleSystem(short namesups)structuralGroup(short namesg)structuralModel(short namesm)structuralParticle(short namesp)structuralRoot(short namesr)visualModel(short namevm)
Examples:
n.t a: matches all atomsn.t sc: matches all side-chain nodesn.t vm: matches all visual models
visibilityFlag
The visibilityFlag attribute (short name vf) matches nodes based on their visibility flag.
Possible values: true or false.
Example:
n.vf true: matches all nodes with a visibility flag set totrue
visible
The visible attribute (short name v) matches nodes that are visible, i.e. the nodes whose visibility flag is true and whose ancestors are visible.
Possible values: none.
Example:
n.v: selects all visible nodes
Atom attributes
Atom attributes are defined in the atom attribute space (short name a). Atom attributes may only match atom nodes.
Possible atom attributes are:
altLocation(short namealt)aminoAcidBackbone(short nameaabb)aromatic(short namear)chain(short namec)chainID(short nameci)customType(short namect)element(short namee)formalCharge(short namefc)hetatm(short namehet)mobile(short namemo)nucleicAcidBackbone(short namenabb)occupancy(short nameoc)partialCharge(short namepc)residueSequenceNumber(short nameresi)resonance(short namereso)serialNumber(short namesn)symbol(short names)temperatureFactor(short nametf)water(short namew)xyz
altLocation
The altLocation attribute (short name alt) matches atoms based on their alternate location.
Possible values: a character.
Example:
a.alt B: matches all atoms with alternate location B
aminoAcidBackbone
The aminoAcidBackbone attribute (short name aabb) matches atoms that belong to an amino-acid backbone.
Possible values: none.
Example:
a.aabb: matches atoms that belong to an amino-acid backbone
aromatic
The aromatic attribute (short name ar) matches atoms that are aromatic.
Possible values: none.
Example:
a.ar: matches aromatic atoms
chain
The chain attribute (short name c) matches atoms that are aromatic.
Possible values: a character.
Example:
a.c A: matches atoms from chain A
chainID
The chainID attribute (short name ci) matches atoms with specific chain IDs.
Possible values: an integer.
Example:
a.ci == 0: matches atoms from chain ID 0a.ci >= 0: matches atoms from chain ID larger than 0a.ci >= 0 and a.ci <=2: matches atoms from chain ID between 0 and 2
customType
The customType attribute (short name ct) matches atoms with specific custom types.
Possible values: an integer.
Example:
a.ct == 0: matches atoms with custom type 0a.ct >= 0: matches atoms with custom type larger than 0a.ct >= 0 and a.ct <=2: matches atoms with custom type between 0 and 2
element
The element attribute (short name e) matches atoms by element.
Possible values: element names.
Example:
a.e Carbon: matches carbon atomsa.e Nitrogen: matches nitrogen atoms
formalCharge
The formalCharge attribute (short name fc) matches atoms with specific formal charges.
Possible values: integer values.
Example:
a.fc >= 1: matches atoms with formal charge larger than 1
hetatm
The hetatm attribute (short name het) matches heteroatoms in protein structures, i.e. atoms whose record type is HETATM in the Protein Data Bank file format.
Possible values: none.
Example:
a.het: matches heteroatoms
mobile
The mobile attribute (short name mo) matches atoms that are mobile.
Possible values: none.
Example:
a.mo: matches mobile atoms
nucleicAcidBackbone
The nucleicAcidBackbone attribute (short name nabb) matches atoms that belong to a nucleic acid backbone.
Possible values: none.
Example:
a.nabb: matches atoms that belong to a nucleic acid backbone
occupancy
The occupancy attribute (short name oc) matches atoms with a specific occupancy.
Possible values: floating-point values.
Example:
a.oc >= 2: matches atoms with occupancy larger than 2
partialCharge
The partialCharge attribute (short name pc) matches atoms with specific partial charges.
Possible values: floating-point values.
Example:
a.pc >= 1.3: matches atoms with partial charge larger than 1.3
residueSequenceNumber
The residueSequenceNumber attribute (short name resi) matches atoms in residues with specific indices.
Possible values: integers.
Example:
a.resi == 12: matches atoms in residue 12a.resi >= 12 and a.resi <=97: matches atoms in residue 12 to 97
resonance
The resonance attribute (short name reso) matches resonant atoms.
Possible values: none.
Example:
a.reso: matches resonant atoms
serialNumber
The serialNumber attribute (short name sn) matches atoms with specific serial numbers.
Possible values: integers.
Example:
a.sn >= 20: matches atoms with serial number larger than 20a.sn >= 20 and a.sn<=897: matches atoms with serial number between 20 and 897
symbol
The symbol attribute (short name s) matches atoms with specific symbols.
Possible values: element symbols.
Example:
a.s C: matches carbon atomsa.s C or a.s H: matches atoms that are carbons or hydrogens
temperatureFactor
The temperatureFactor attribute (short name tf) matches atoms with specific temperature factors.
Possible values: floating-point values.
Example:
a.tf > 2: matches atoms with a temperature factor strictly larger than 2
water
The water attribute (short name w) matches water atoms.
Possible values: none.
Example:
a.w: matches water atoms
x
The x attribute matches atoms with specific x coordinates
Possible values: floating-point values with length units.
Example:
a.x >= 1.0 A: matches atoms whose x coordinate is larger than 1.0 A
Possible units for lengths are:
micrometer(short nameum)nanometer(short namenm)angstrom(short nameA)picometer(short namepm)femtometer(short namefm)
y
The y attribute matches atoms with specific y coordinates
Possible values: floating-point values with length units.
Example:
a.y >= 1.0 A: matches atoms whose y coordinate is larger than 1.0 A
Possible units for lengths are:
micrometer(short nameum)nanometer(short namenm)angstrom(short nameA)picometer(short namepm)femtometer(short namefm)
z
The z attribute matches atoms with specific z coordinates
Possible values: floating-point values with length units.
Example:
a.z >= 1.0 A: matches atoms whose z coordinate is larger than 1.0 A
Possible units for lengths are:
micrometer(short nameum)nanometer(short namenm)angstrom(short nameA)picometer(short namepm)femtometer(short namefm)
Bond attributes
Bond attributes are defined in the bond attribute space (short name b). Bond attributes may only match bond nodes.
Possible bond attributes are:
customType(short namect)order(short nameo)
customType
The customType attribute (short name ct) matches bonds with specific custom types.
Possible values: an integer.
Example:
b.ct == 0: matches bonds with custom type 0b.ct >= 0: matches bonds with custom type larger than 0b.ct >= 0 and b.ct <=2: matches bonds with custom type between 0 and 2
order
The order attribute (short name o) matches bonds with specific orders.
Possible values: floating-point values.
Example:
b.o >= 2: matches all bonds with order larger than 2
Residue attributes
Residue attributes are defined in the residue attribute space (short name r). Residue attributes may only match residue nodes.
Possible residue attributes are:
secondaryStructure(short namess)type(short namet)
secondaryStructure
The secondaryStructure attribute (short name ss) matches residues with specific secondary structures.
Possible values:
alpha(short namea)beta(short nameb)helix(short nameh) (same matches asalpha)strand(short names) (same matches asstrand)
Example:
r.ss h: matches all residues in alpha helices
type
The type attribute (short name t) matches residues with specific types.
Possible values:
ACGUIDADCDGDTDIALAARGASPASNVALHISGLYGLUGLNILELEULYSMETPROSERTYRTHRTRPPHECYSASXGLXXLEXAASECPYL
Example:
r.t ALA: matches all alanines
Specification by name
Nodes may be specified by names, i.e. strings enclosed with quotes. In the SAMSON data graph, nodes which may have a custom name are:
- atoms
- backbones
- cameras
- chains
- conformations
- dynamical models
- groups
- interaction models
- labels
- folders
- molecules
- property models
- residues
- segments
- side-chains
- simulators
- structural groups
- structural models
- visual models
Note that bond names are formed from the names of the atoms they bond, and cannot be searched by name.
As noted above, pressing the Tab key in the search box when a string is being entered (with the quote sign, to indicate a name) lists all nodes whose name begins with the entered string. For example, entering "ALA and pressing the Tab key may yield e.g:
"ALA 22 Backbone""ALA 22 Side-chain""ALA 22""ALA 28 Backbone""ALA 28 Side-chain""ALA 28"...
Possible values: a string enclosed with quotes.
Examples:
"Folder 1""CA""Group 1""1EQZ""ALA 28""Nanotube""Simulator 1"
Specification by symbols and element names
In order to efficiently match atoms of a given element type, symbols and element names are valid NSL expressions.
Possible values: atomic symbols and element names (capitalized)
Examples:
Carbon: matches all carbon atomsC: matches all carbon atomsCa: matches all calcium atomsH: matches all hydrogen atoms
Operations on sets
Logical operators
Logical operators may be used to perform operations on sets.
Possible values: and, not, or and xor (exclusive or).
Examples:
a.sn >= 20 and a.sn <= 897: matches atoms with serial number between 20 and 897a.sn <= 20 or a.sn >=40: matches atoms with serial number smaller than 20 or larger than 40n.t r and not r.t CYS: matches residue nodes that are not cysteinsa.sn >= 20 xor a.oc >= 0.5: matches atoms which either have a serial number larger than 20, or have an occupancy larger than 0.5, but not those that satisfy both conditions
Note that, without proper care, the not condition might produce surprising results. For example, not r.t CYS does not return the list of residues that are not cysteins. Indeed, a folder node is also a node which is not a residue node that's a cystein. If only residues nodes should be returned, a proper query would be n.t r and not r.t CYS.
Topology operators
Containement operators may be used to specify inclusion in or out of a set.
Possible values: in and out of.
Examples:
n.t a in "2AZ8": matches atoms in "2AZ8"n.t a in r.t CYS: matches atoms that belong to cysteinsH in r.t ARG: matches hydrogens that belong to argininsH in n.h: matches hidden hydrogensn.t a out of r.t PRO: matches atoms that do not belong to prolines
Proximity operators
Proximity operators may be used to select nodes based on distances.
Possible values: within {distance} of and beyond {distance} of.
Examples:
C within 5A of "GLN 2": matches carbons within 5 angstrom of "GLN 2"n.t a beyond 5A of "2AZ8-IA": matches atoms beyond 5 angstrom of "2AZ8-IA"
Possible units for distances are:
micrometer(short nameum)nanometer(short namenm)angstrom(short nameA)picometer(short namepm)femtometer(short namefm)
Note that, without proper care, the within condition might produce surprising results. For example, * within 5A of "GLN 2" will select the document node, since some atoms in the document are within 5 angstrom of "GLN 2": the atoms that belong to "GLN 2". If only atom nodes should be returned, a proper query would be n.t a within 5A of "GLN 2". Note that this latter query would also return atoms that belong to "GLN 2". If only atoms outside "GLN 2" should be returned, then a proper query would be n.t a within 5A of "GLN 2" out of "GLN 2".
Managing groups
As noted above, selections may be saved as groups in the data graph. For example, "Interface" = ((n.t a in "A") w 4A of "B") or ((n.t a in "B") w 4A of "A") creates a group containing atoms at the interface between "A" and "B". Precisely, a group node called "Interface" is added to the document.
Note that "node group" is a specific node type. As a result, the query "Interface" will return the group node itself, and not the nodes forming the group.
Reference
Attributes
Node attributes
| Attribute name | Short name | Possible values | Examples |
hidden | h | n.h | |
selected | s | n.s | |
selectionFlag | sf | true or false | n.sf true |
type | t | See below | n.t a |
visibilityFlag | vf | true or false | n.vf false |
visible | v | n.v |
For the type attribute, possible values are
atom(short namea)camera(short nameca)backbone(short namebb)bond(short nameb)chain(short namec)conformation(short nameco)document(short named)dynamicalModelParticleSystem(short namedmps)interactionModelParticleSystem(short nameimps)label(short namela)folder(short namef)molecule(short namem)nodeGroup(short nameng)propertyModel(short namepm)pseudoatom(short namepa)residue(short namer)segment(short names)sidechain(short namesc)simulatorParticleSystem(short namesps)stateUpdaterParticleSystem(short namesups)structuralGroup(short namesg)structuralModel(short namesm)structuralParticle(short namesp)structuralRoot(short namesr)visualModel(short namevm)

