In this tutorial, we will create an importer for the following XYZ format: The 1st line: the number of atoms in the file. The 2nd line: arbitraty header line. Next lines contain information on atoms in the following format (coordinates are in angstroms): element_symbol x-coordinate y-coordinate z-coordinate.
Example of the format:
Note: SAMSON already has an XYZ Importer which is installed by default, that's why we are actually going to read files with a exyz extension (e for element), even though the contents will still correspond to the xyz format. Therefore, to test this importer, you can export (File menu > Save as) any system from SAMSON in the XYZ format and change the extension from .xyz to .exyz.
The source code for an Element created in this tutorial can be found at https://github.com/1A-OneAngstrom/SAMSON-Developer-Tutorials.
Generating an Element
We will start by creating a new SAMSON Element called EXYZ thanks to the SAMSON Element generator (please, refer to the SAMSON Element generator tutorial for a reminder on how to use it):
- launch SAMSON and run the SAMSON Element generator;
- specify the path to a folder where you want to develop your SAMSON Elements;
- name the SAMSON Element as EXYZ and add some description;
- add an Importer class (called
SEEXYZImporter); - generate the SAMSON Element.
The SAMSON Element generator generates an importer class (derived from the SBIFileImporter class) and a GUI class that implements the user interface of the importer and is derived from the SBGFileImporter class.
Parsing a file
In SAMSON, an importer is created by programming three functions in the importer class:
- getExtension - specifies the extension of a file;
- getFilter - specifies a filter in Qt format;
- importFromFile - a function invoked by SAMSON to import a file with a suitable extension.
Let's open the SEEXYZImporter.cpp file and implement these functions.
Modify the getExtension function to return the extension of the importer (exyz) as follows:
Modify the getFilter function to return the filter of the importer as follows (it should be in the Qt format):
We now have to code the importFromFile function to read a molecule from a exyz file and create a structural model. To achieve this, we are going to use the SBIFileReader class to read a file, the SAMSON class to manage the undo mechanism and access the active document, and several structural classes. For that, include the following headers in the file:
Let's now code inside the importFromFile function:
We first begin by checking if the file exists and if it is readable:
Now we open the file and store all its lines in a vector of strings:
Then, we read the number of atoms in the file:
We create a new structural model:
And we read and create atoms that we add to the structural model:
After adding atoms in the structural model we can create covalent bonds between these atoms:
Now we can add the structural model to the document:
SAMSON::beginHolding and SAMSON::endHolding functions turn the undo system on and off, respectively. The SAMSON::hold function tells SAMSON to acquire ownership of the structural model and its contents. Line structuralModel->create(); sets the structural model to a created state (i.e. not erased). Then we implement the expected behavior of an importer, which is to import to the active document by default, unless a folder (preferedFolder) has been specified.
Finally, we signal that we had no issue with the file (in this version of the importer, we do not deal with potential reading errors):
That's it, we have parsed the file, filled in a structural model, and added it into a SAMSON document.
If you want for the Importer to get some parameters/options from the user, please follow to the section Getting parameters, if not then you can simply disable the options window by replacing the constructor and destructor of SEEXYZImporter as follows:
Getting parameters
For some file formats it might be necessary for user to specify some parameters/options for an importer. Parameters can be received either from the GUI of an Importer or through the importFromFile function itself if it was invoked from the code. In this section we will show how to implement this.
Let's consider the possibility for the EXYZ Importer to import with or without creation of covalent bonds.
Open the SEEXYZImporterGUI.ui file in QtCreator. Remove the label with a tip and add a QCheckBox with a text as follows:
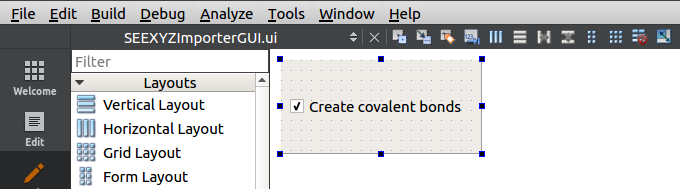
Give a meaningful name to the check box: checkBoxCreateCovalentBonds
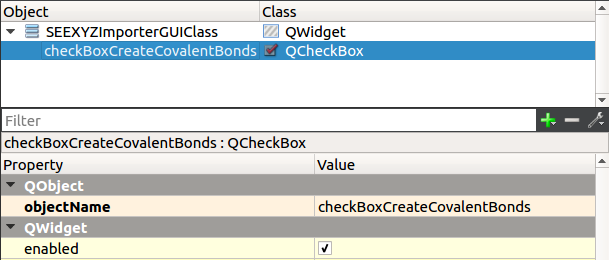
We want to save the GUI state (the state of the check box) from one session to the next. For that, we need to implement saveSettings and loadSettings functions. The saveSettings function is invoked when a SAMSON Element is being destructed. The loadSettings function is invoked in the constructor of a SAMSON Element.
Here we save the current state of the check box in the variable checkBoxCreateCovalentBonds which will be stored thanks to the QSettings functionality.
Here we set the state of the check box to the saved one.
It is a good practice to separate GUI and non-GUI functionalities, e.g. do not access directly GUI elements from an another class. That is why we add the following function in the SEEXYZImporterGUI class (SEEXYZImporterGUI.hpp file):
This function will be used in the SEEXYZImporter to get parameters from the GUI class. Implement it as follows.
Add the following private variable in the SEEXYZImporter class (SEEXYZImporter.hpp file):
Add the following functions in the SEEXYZImporter class (SEEXYZImporter.hpp file):
Implement the initializeParameters function that initializes parameters for the parser.
In this code, if no parameters has been specified (the parameters list is empty or has the wrong size) we take them from the GUI form, else we parse the parameters list for which we implement the parseParameters function as follows:
Here we go through the parameters list and read the parameters.
Now, invoke initializeParameters function in the beginning of the importFromFile function:
This allows both taking care of parameters specified through GUI or through direct invocation of the SAMSON::importFromFile function in the code.

