Hi everyone,
In this post, I will show you how to use the Ligand Unbinding Pathway Search app for finding possible ligand unbinding pathways from a protein. Let us start with a list of requirements, and then a tutorial on how to use the app.
Requirements
- SAMSON 0.7.0
- The Ligand Unbinding Pathway Search app
- The GROMACS force field interaction model
- The FIRE state updater
Before starting the tutorial, I would like to ask you to download this file (LigandUnbindingPathSearch) which contains necessary files for this tutorial. After extracting it, you will have a structural model (1pv7_ligand_complex.sam) and the topology files for energy evaluation with Gromacs (1pv7_ligand.itp, 1pv7_ligand_complex_topol.top, 1pv7_porse.itp).
Tutorial
In this tutorial, we will apply the ART-RRT method for finding ligand unbinding pathway from a protein. The ART-RRT method is a combination of T-RRT method and ARAP modeling method. The T-RRT method is used for finding the possible pathways and the ARAP modeling method is for generating possible ligand motions. Moreover, a constrained minimization is used for adapting the motion of the protein with the ligand. Let us begin.
Loading the input model
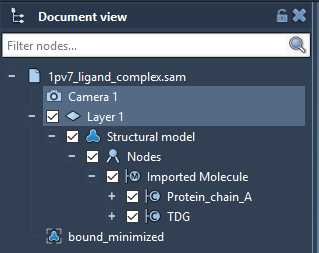
Run SAMSON 0.7.0. Load 1pv7_ligand_complex.sam. It will open a structural model of Lactose permease with its ligand Thiodigalactosid. In the Document View (if you can not find it, try showing it under Window in the toolbar), you can see the Protein under the name Protein_chain_A and the ligand under the name TDG as shown in the picture below. You also see 1 conformation named bound_minimized which is the minimized conformation of the protein-ligand complex.
Creating Secondary Structure
We will now create a secondary structure for the protein by right-clicking on Protein_chain_A in the Document View, choose Add> Visual Model. In the Add visual model window, set the parameters as shown in the following figure.
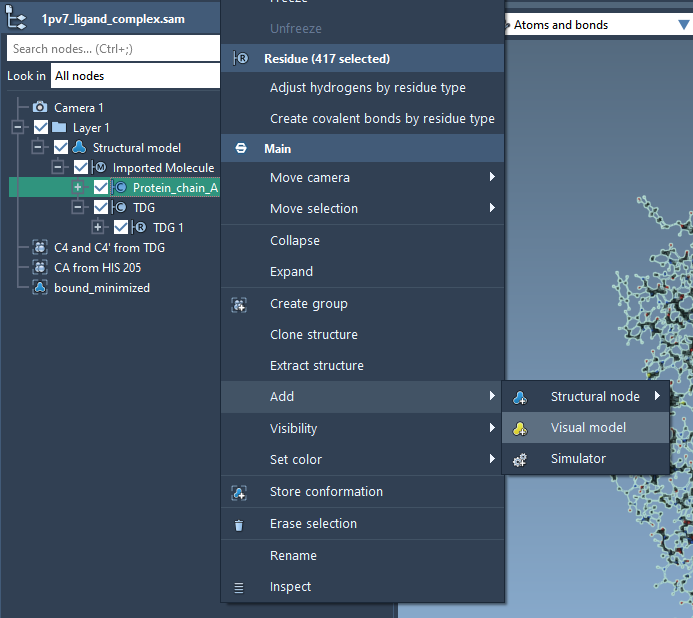
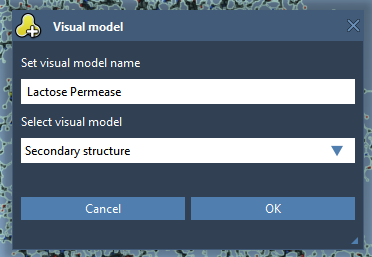
Next, we hide the ball-and-stick representation of the protein by unchecking the box next to the Protein_chain_A inside the Document View for more visibility.
Executing Ligand Unbinding Pathway Search app
Now open the Ligand Unbinding Pathway Search app by clicking on its icon ![]() or you can find its name under App in the toolbar. In the app’s window, you will see two tabs where tab Settings is for setting up the parameters and tab Results is for collecting the results.
or you can find its name under App in the toolbar. In the app’s window, you will see two tabs where tab Settings is for setting up the parameters and tab Results is for collecting the results.
Setting up interaction and state updater models for energy evaluation
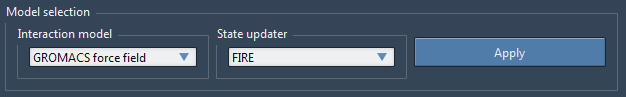
Let us stay inside the tab Settings. Under Model Selection, select GROMACS force field in the Interaction model drop-down list, and FIRE in the State updater drop-down list, then click Apply (see the figure below). This will apply the interaction model and state updater on all the atoms currently in the SAMSON view.
A window pops up asking if you want to continue, click Yes. The GROMACS force field setup will then ask for the topology file for the protein-ligand model. Click Browse, then choose to open the file 1pv7_ligand_complex_topol.top provided with the structural model file. Keep the rest of the information in the window intact (see the figure below) then click OK.


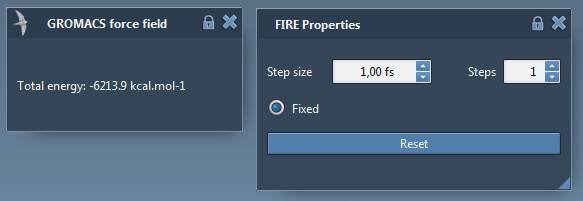
You will see GROMACS force field window popping up for showing the potential energy of the current state of the system, and FIRE Properties window for showing the parameters from the state updater. Set the parameters for FIRE as below
Defining the ligand-protein bound state
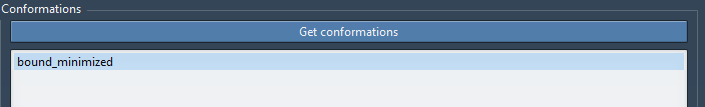
Under Conformations, click Get conformations to obtain all the current conformations in the Document View. Then, select bound_minimized in the list as shown below. It will consider the bound_minimized conformation as the start state for growing the T-RRT tree.
Assigning the atom types and defining the sampling box
Now check the box DoF editor to start assigning atom types.
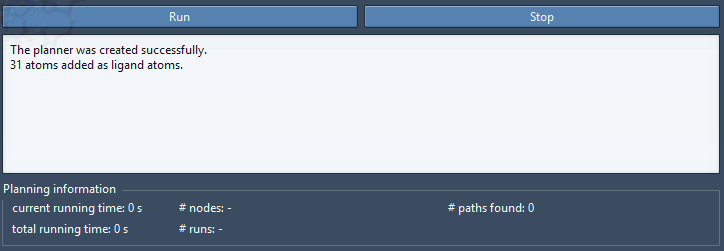
Select TDG in the Document view to select all the ligand atoms. Click Add ligand atoms button to tell the app that the selected atoms are ligand atoms. The rest of the atoms will be implicitly considered as protein atoms. A new line appears in the text box at the bottom of the app saying that “31 atoms added as ligand atoms.” as shown in the figure below. This text box is for additional information.
Next, expand the TDG molecule (the ligand) in the Document View up to its SideChain, select C4 and C4’ atoms (Ctrl + left-click for multiple selection), then press Add active ARAP atoms button in the app to use these atoms for controling the ligand motion. The motions of the rest of the ligand atoms (implicitly considered as passive ARAP atoms) follow these two atoms by the ARAP modeling algorithm.
Finally, select the CA atom in the Backbone of HIS 205 residue of the protein and click Add fixed atoms button in the app to fix its position during the search. This is just a safety measure so that the protein molecule does not drift along with the ligand.
If you are not satisfied with the atom type assignment, click the Reset button the start over.
During the atom type assignment, you can notice that another visual model is created for showing the atom type (blue for passive ARAP atoms, green for active ARAP atoms, and red for fixed atoms).
We will now design the sampling box for the active ARAP atoms by checking the box Show sampling box for active ARAP atoms. Set the box dimension as such:
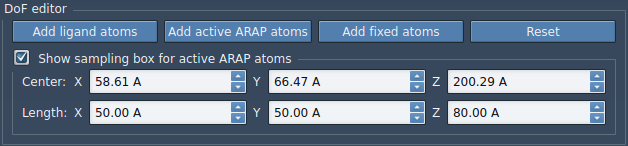
A green box will show up in SAMSON for visualizing this box. This box defines the same sampling region for the chosen active ARAP atoms. This box dimension bias the ligand motion toward the periplasmic side of the protein.
Defining the parameters
Under Parameters, set them as in the following figure:
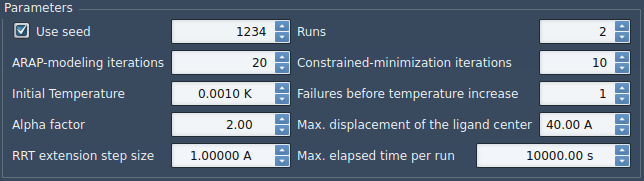
- Use seed box is checked and value is 1234: we use a seed of 1234 for the planner and after each run the seed value is incremented by 1. If the box is unchecked, a random seed is used.
- Runs = 2, we run the method 2 times to extract a maximum of 2 paths.
- ARAP-modeling iterations = 20: the number of iterations for solving ARAP modeling.
- Constrained-minimization iterations = 10: apply 10 steps of constrained minimization with FIRE each time a new state is sampled to minimize it.
- Initial Temperature = 0.001 K, Alpha factor = 2, Failures before temperature increase = 1 : parameters for T-RRT.
- Maximum displacement of the ligand center = 40 A: each run is stopped as soon as the center of the ligand is displaced by 40 angstrom from its original position.
- RRT extension step size = 1 A: the planner extension step size (or edge length in the RRT tree) is 1 angstrom. Note that this is only the prescribed edge length. The final edge lengths can deviate a bit from this value.
- Maximum elapsed time per run: each run is stopped as soon as the elapsed time reaches this value.
Running, Pausing, or Stopping the planner
Click the Run button to start the planner. The searching process can be paused by clicking the Pause button and resumed after that by clicking on the Resume button (these buttons are located at the same place as the Run button while the planner is running). To stop the process, click the Stop button.
During the execution, you can observe the time elapsed for the current run (Current running time), the time elapsed for all of the runs (Total running time), the number of nodes of the tree in the current run (Nodes), the run number (Run), the number of paths found (Paths found), under the Planning information as shown in the figure below.

Obtaining the results
As soon as a path is found, it is added to a list in the Results tab. For example, the following figure shows two paths found.
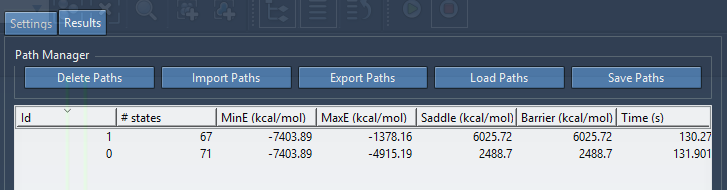
Each path contains the following field
- id: path id
- # states: Number of conformations (or states) in the path.
- MinE (kcal/mol): Minimum energy of the path conformations.
- MaxE (kcal/mol): Maximum energy of the path conformations.
- Saddle (kcal/mol): Difference between MaxE and MinE.
- Barrier (kcal/mol): Difference between MaxE and First.
- Time (s): Time elapsed for searching this path.
- First (kcal/mol): Energy of the first conformation in the path.
- Last (kcal/mol): Energy of the last conformation in the path.
- Remarks: Comments on the path (editable).
- Color: Color of the energy curve for this path.
Viewing conformation energies along the path
To view the energy curve of the path, select a path by clicking on it in the path list. To plot several energy curves, select several paths (by Ctrl + left-clicking for multi-selection).
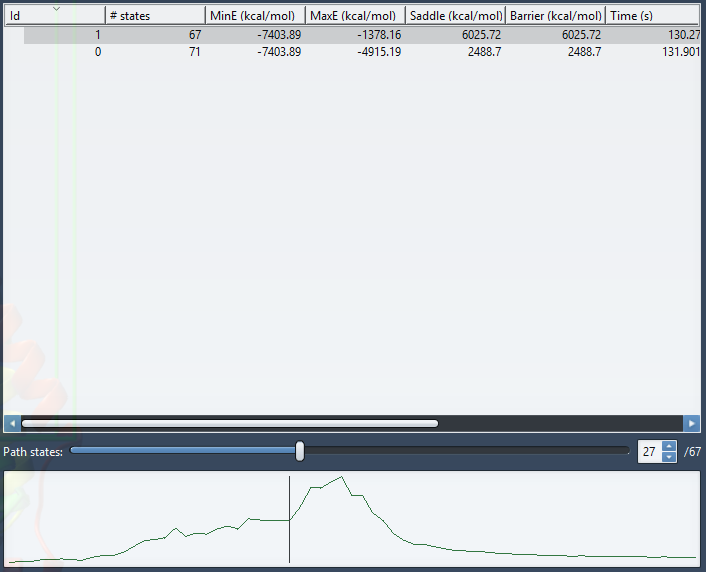
After selecting a path, you can move the slider to select a particular conformation in the path as shown in the above figure. The corresponding conformation is reflected in the structural model in SAMSON and its corresponding energy is shown in the GROMACS force field window.
Exporting path energies or the path list content
Path energy values and the path list content can also be copied to the clipboard by first selecting the paths you are interested in in the list, then right-clicking as shown in the figure below
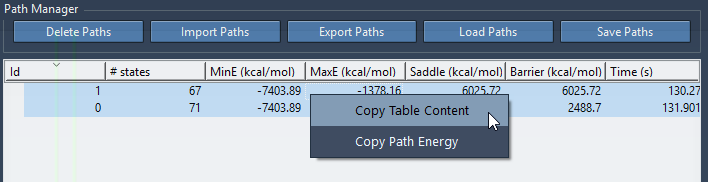
The Path Manager offers other functionalities on paths:
- Delete Paths button: delete selected paths from the list.
- Import Paths button: select a list of SAMSON conformations and then click this button to import them as a path in the list. Note that to be able to do this, you must apply a model selection under the Settings tab first.
- Export Paths button: create SAMSON conformations for the selected paths in the list.
- Load Paths button: load paths from a cpath file. Note that to be able to do this, you must apply a model selection under the Settings tab first.
- Save Paths button: save all the selected path in the list in a cpath file.
Next steps
Creating pathlines
Check the Pathlines tutorial to learn about how to create pathlines, for example to visualize the movement of the center of mass of a ligand.
Improving paths with P-NEB
The resulting paths can be significantly improved with the help of the parallel Nudged Elastic Band (NEB) method implemented in the P-NEB app. Please check out the P-NEB tutorial for more information.
Exporting atoms trajectories along paths
The resulting paths can be saved in the .sam and .samx formats. If you want to export only trajectories of some atoms along the path, you can use the Export Along Paths app. Please refer to the tutorial on how to use the Export Along Paths app for more information.
If you have any questions or feedback, please use the SAMSON forum.





