To get SAMSON, sign up on SAMSON Connect (it’s free!) and download it now!
In line with our vision of a platform democratizing access to molecular modeling, SAMSON 2020 significantly improves the user experience and brings numerous, game-changing functionalities.
As easy as “ABC”
We’ve wanted to offer this in SAMSON for years, and it’s finally here: a powerful yet easy-to-use Molecular builder.
A is for “Atoms” and “Assets”
Of course, we made it possible to build using individual atoms:
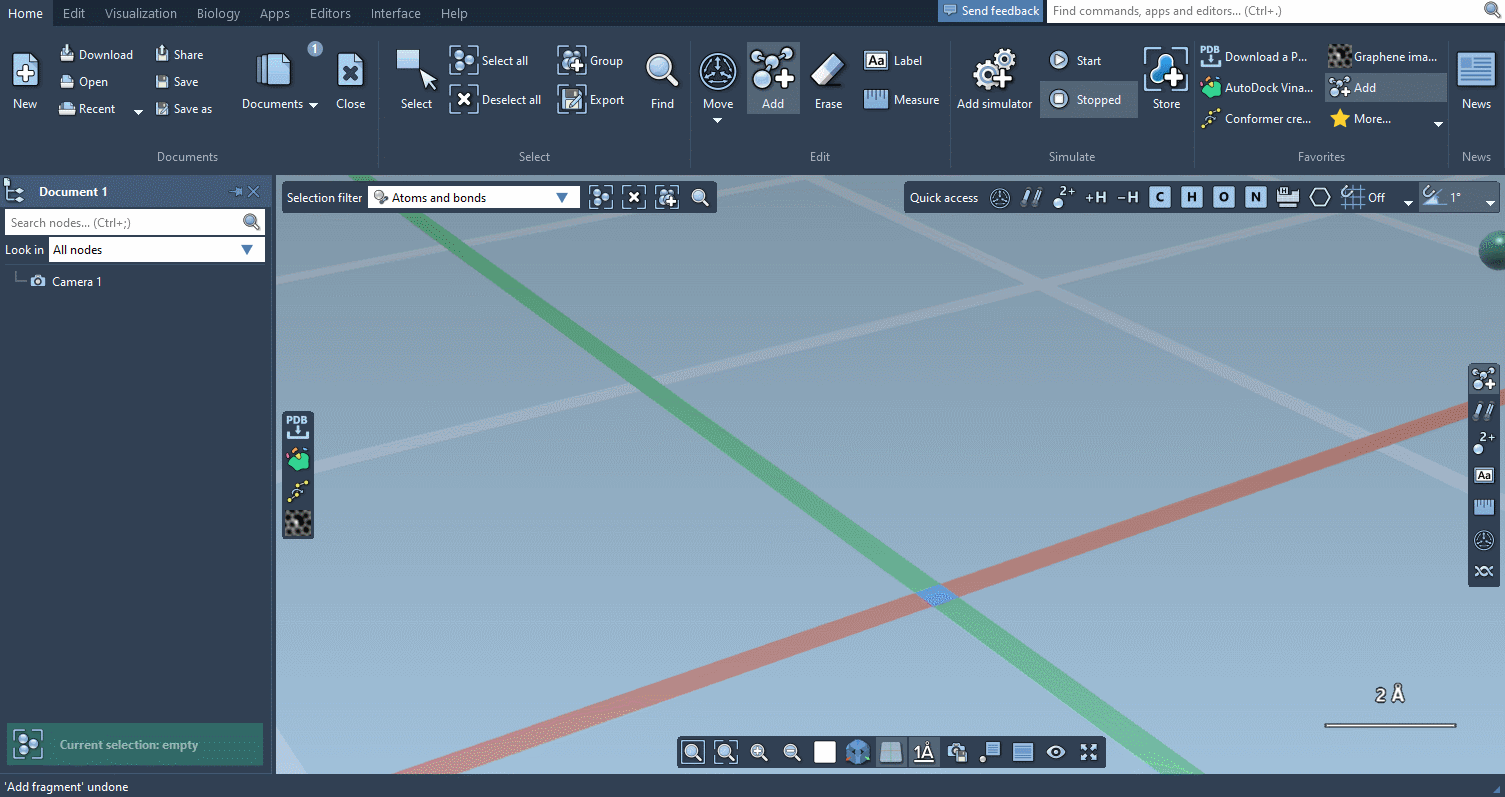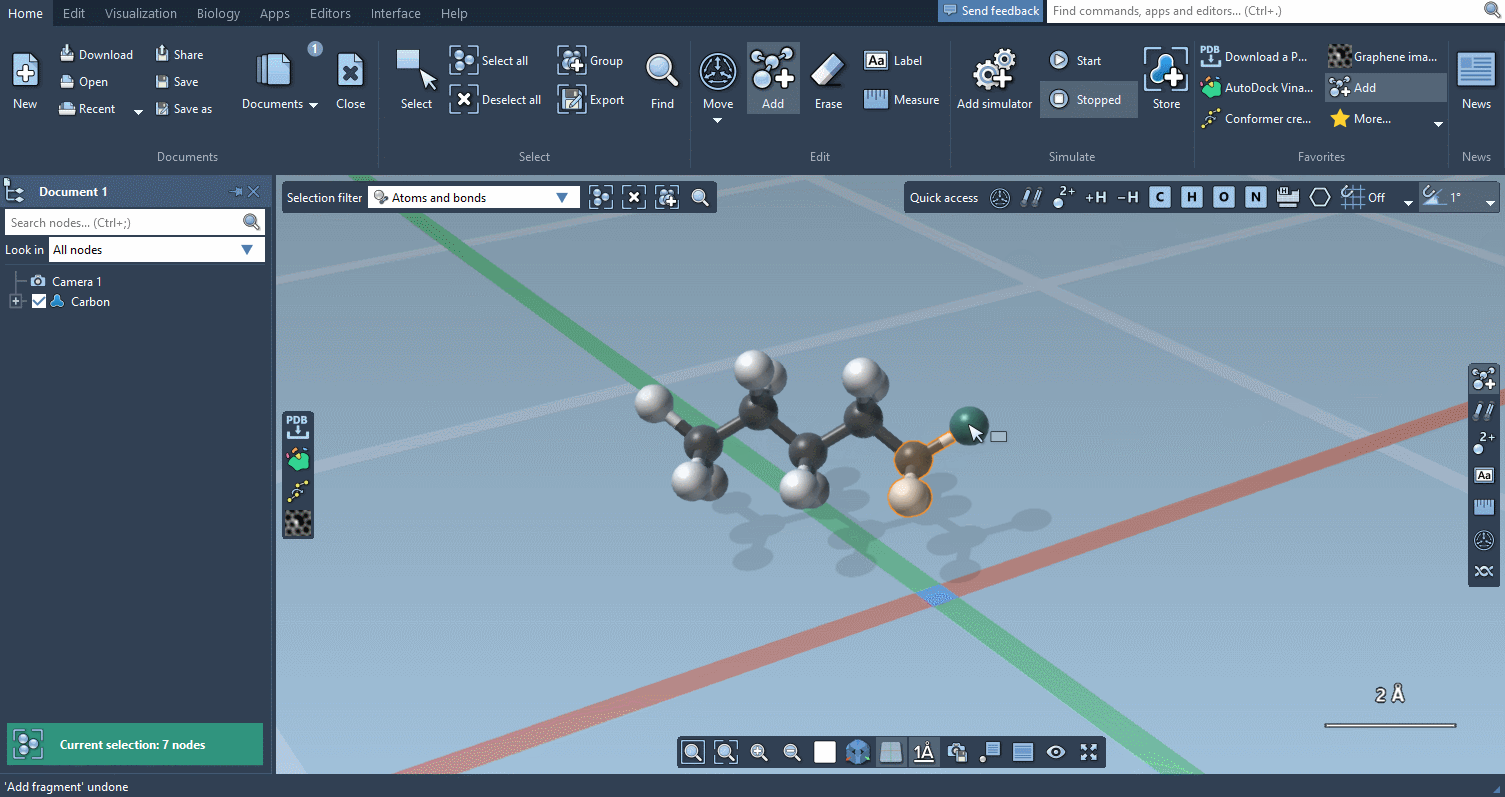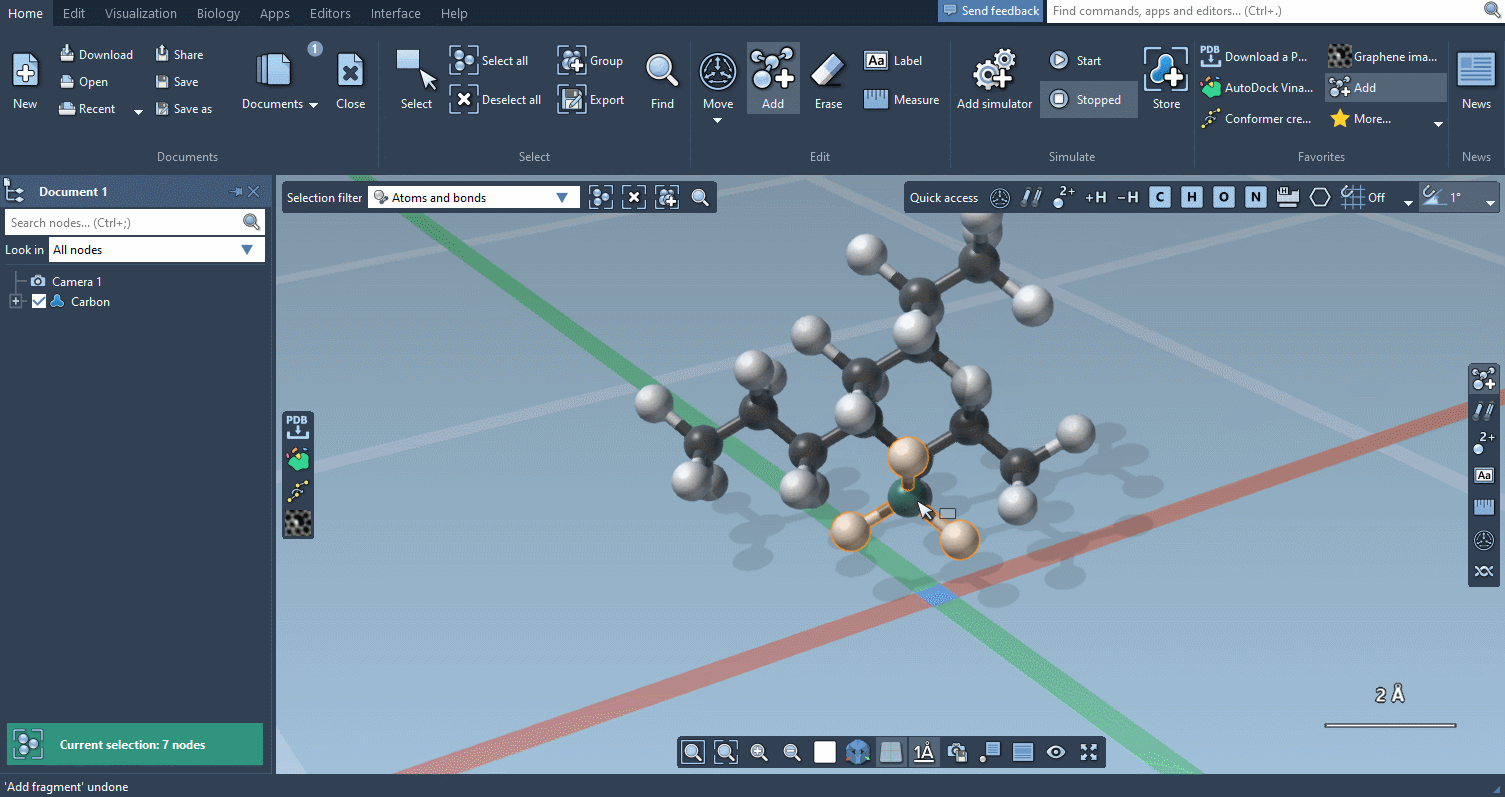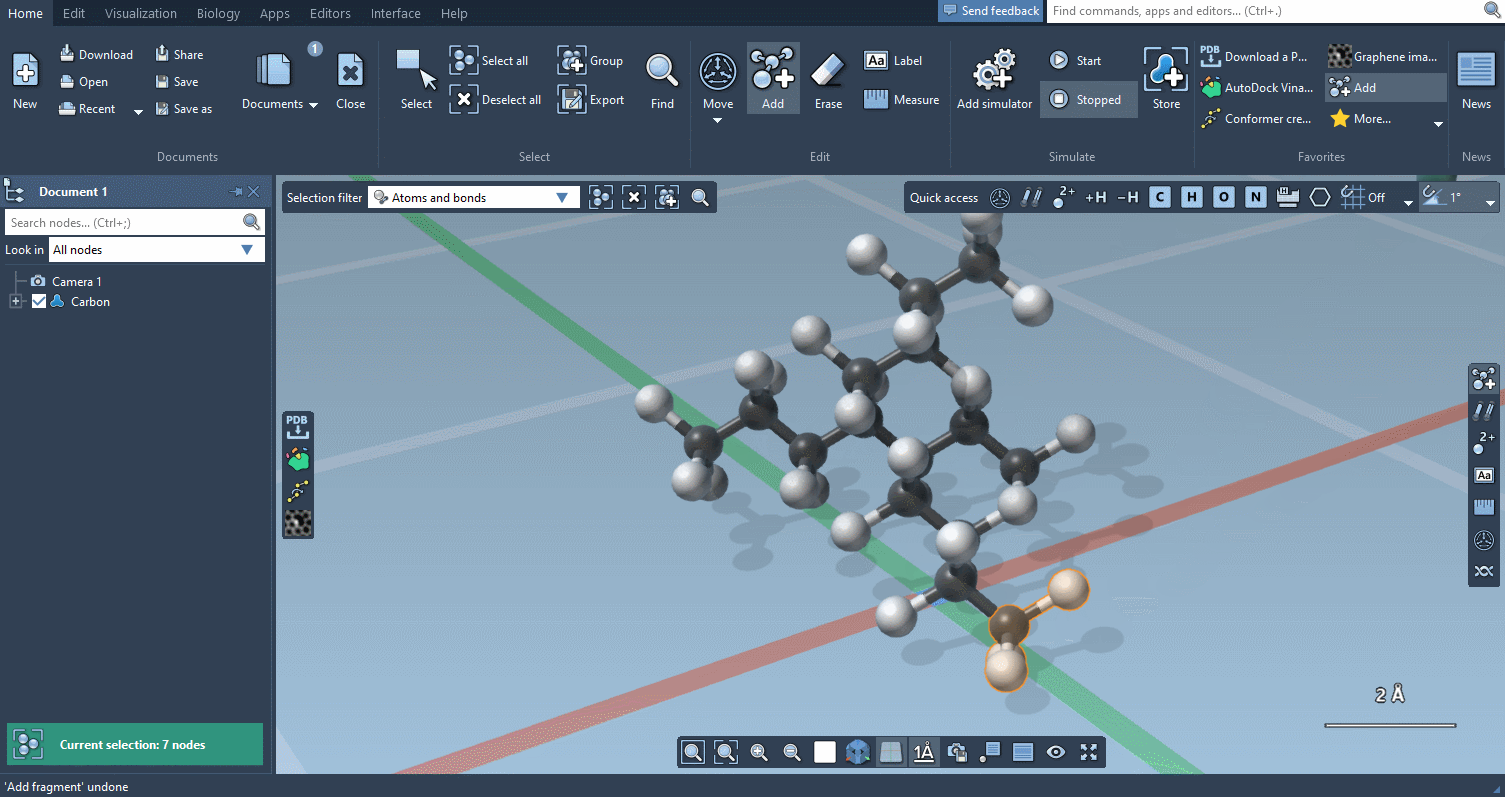
You can choose which atoms you add using Quick commands…
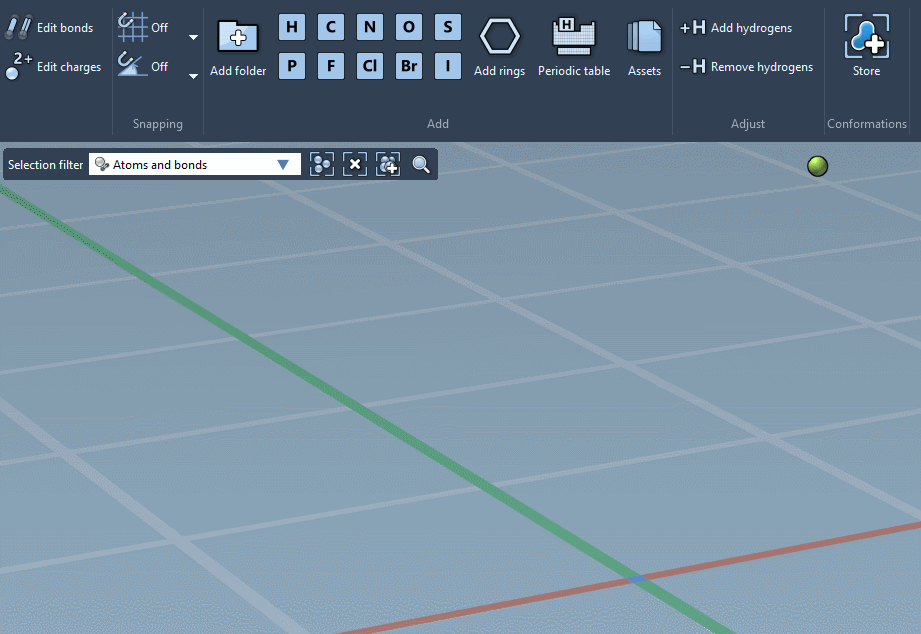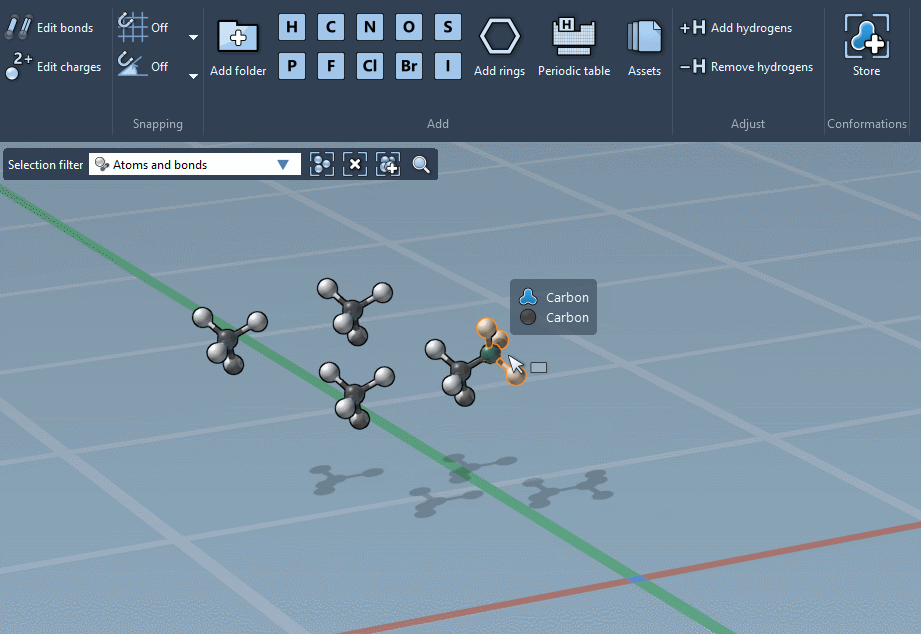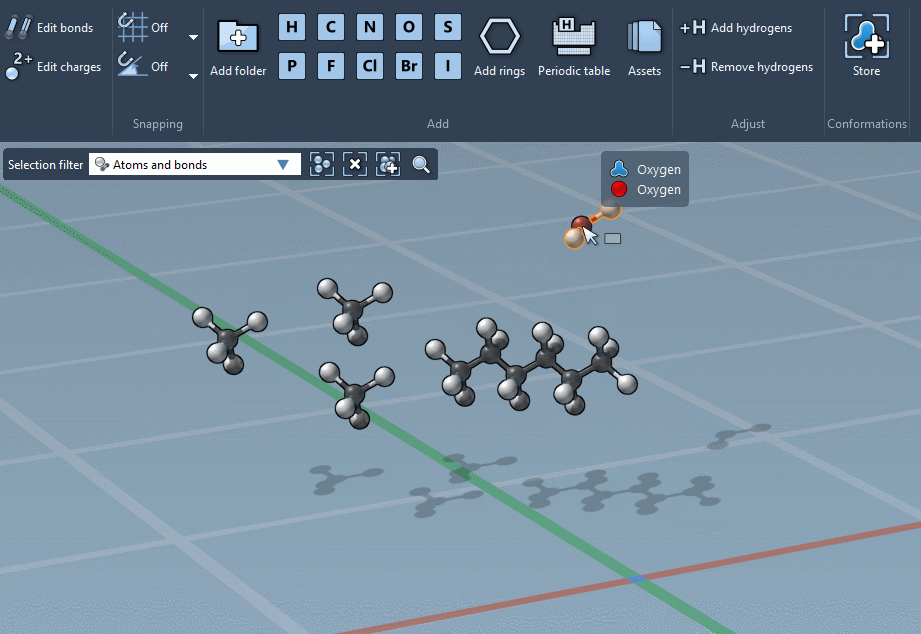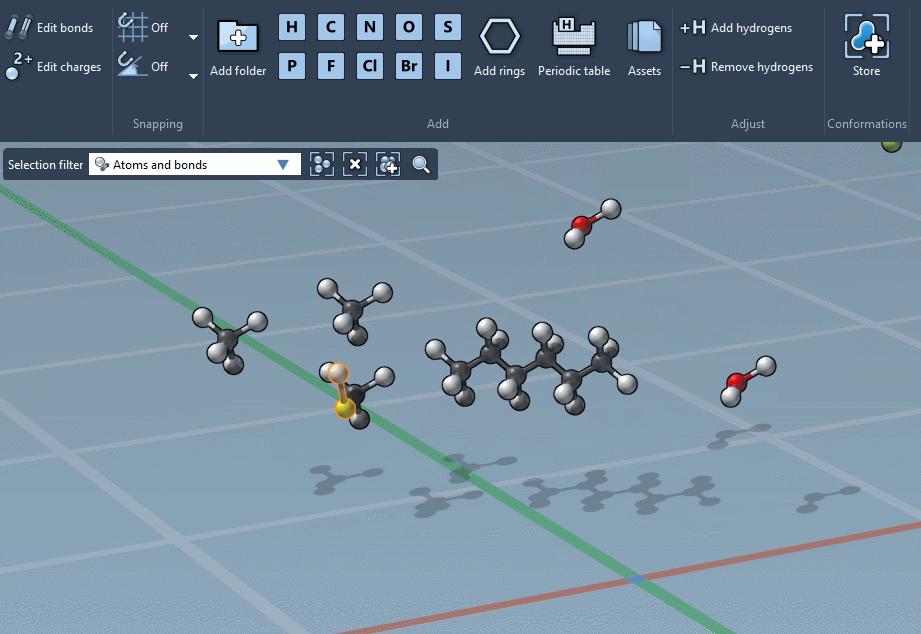
… or using the Periodic table…
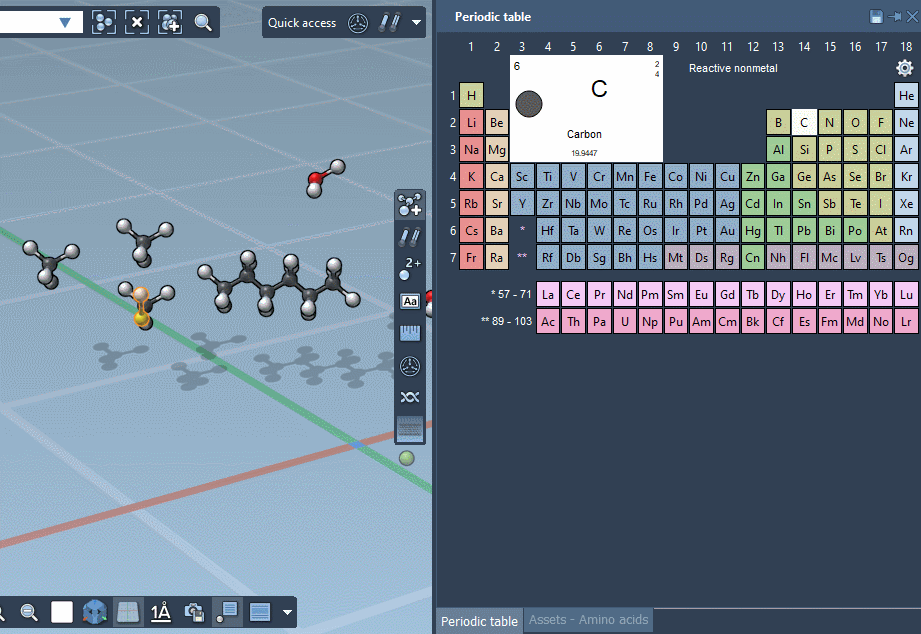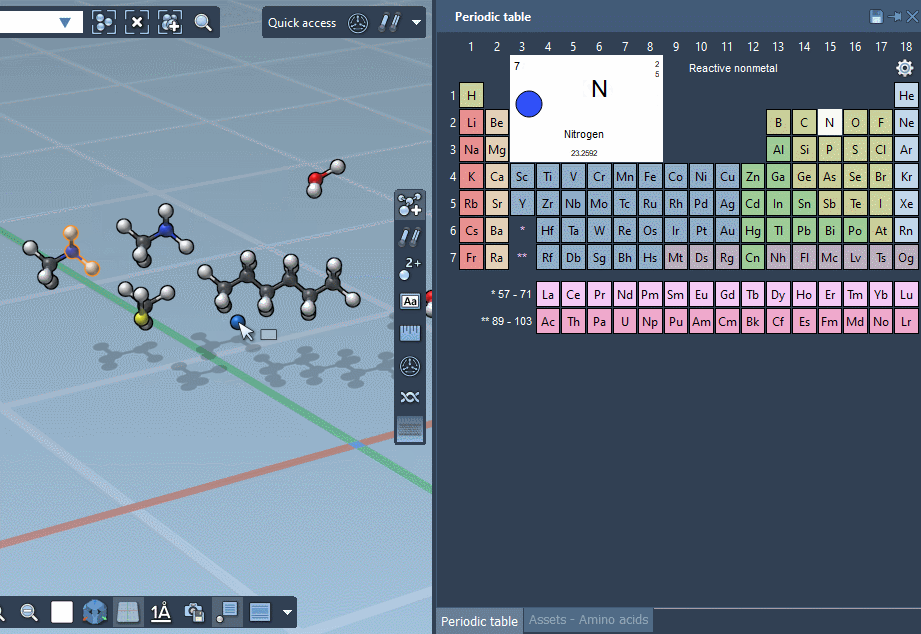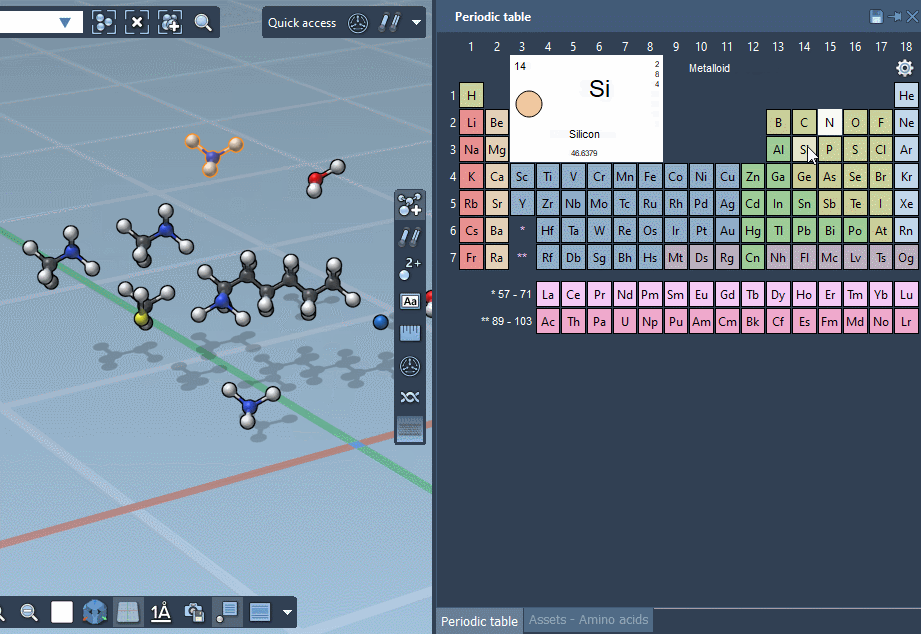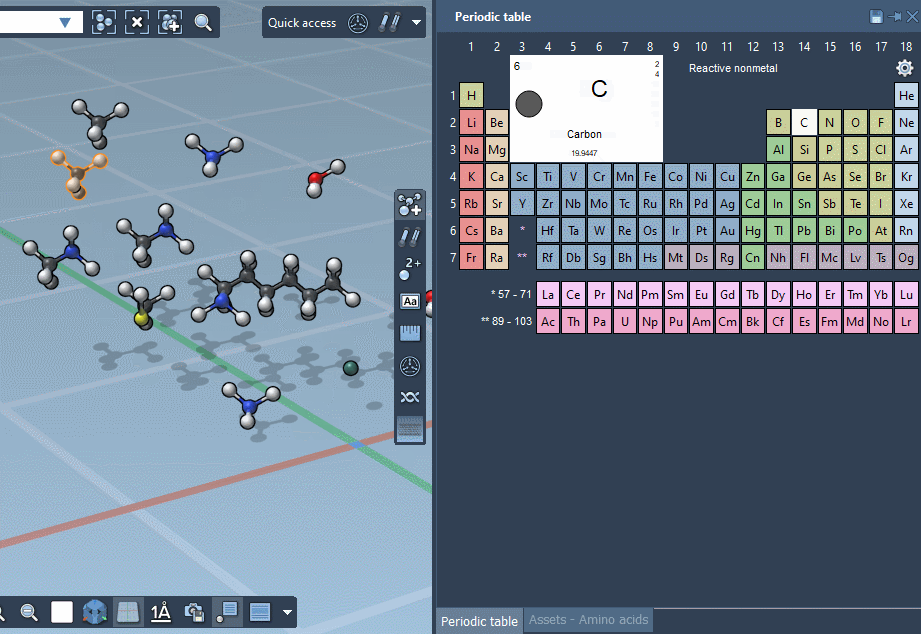
… but we’re excited to announce that SAMSON lets you build by assembling assets!
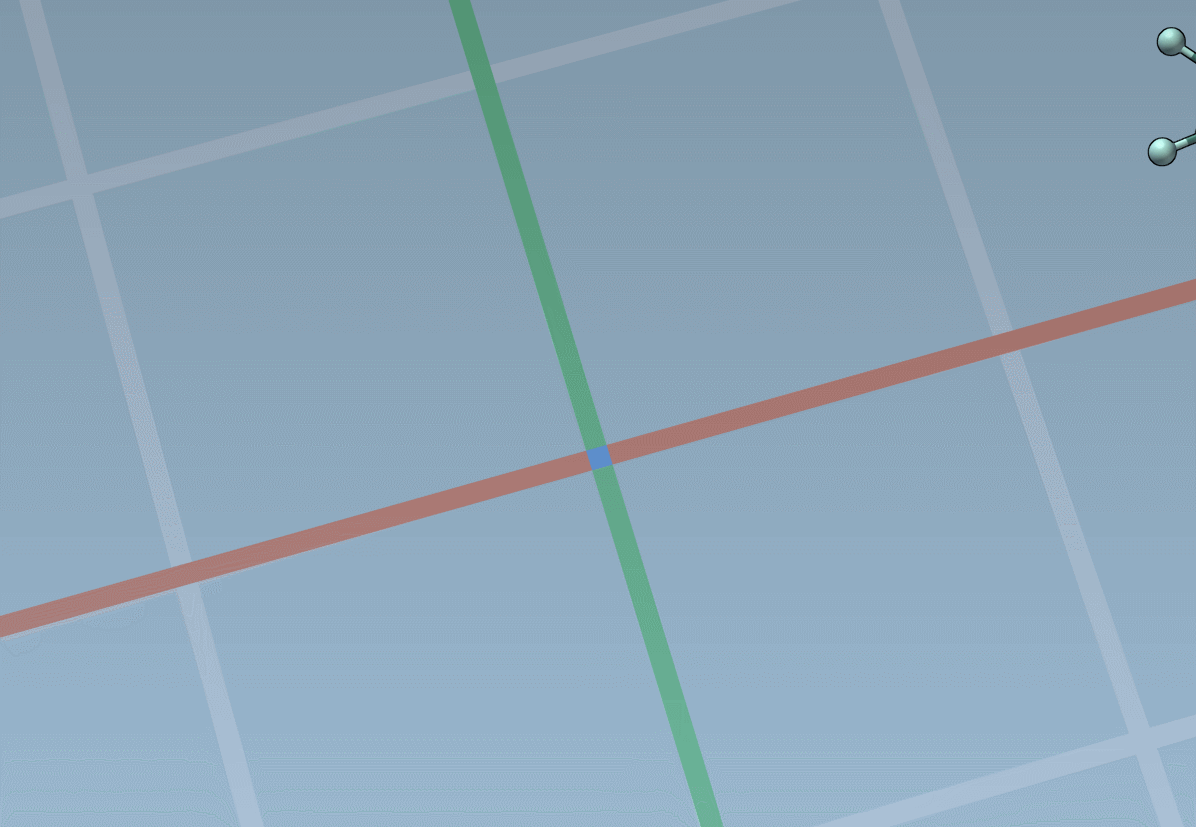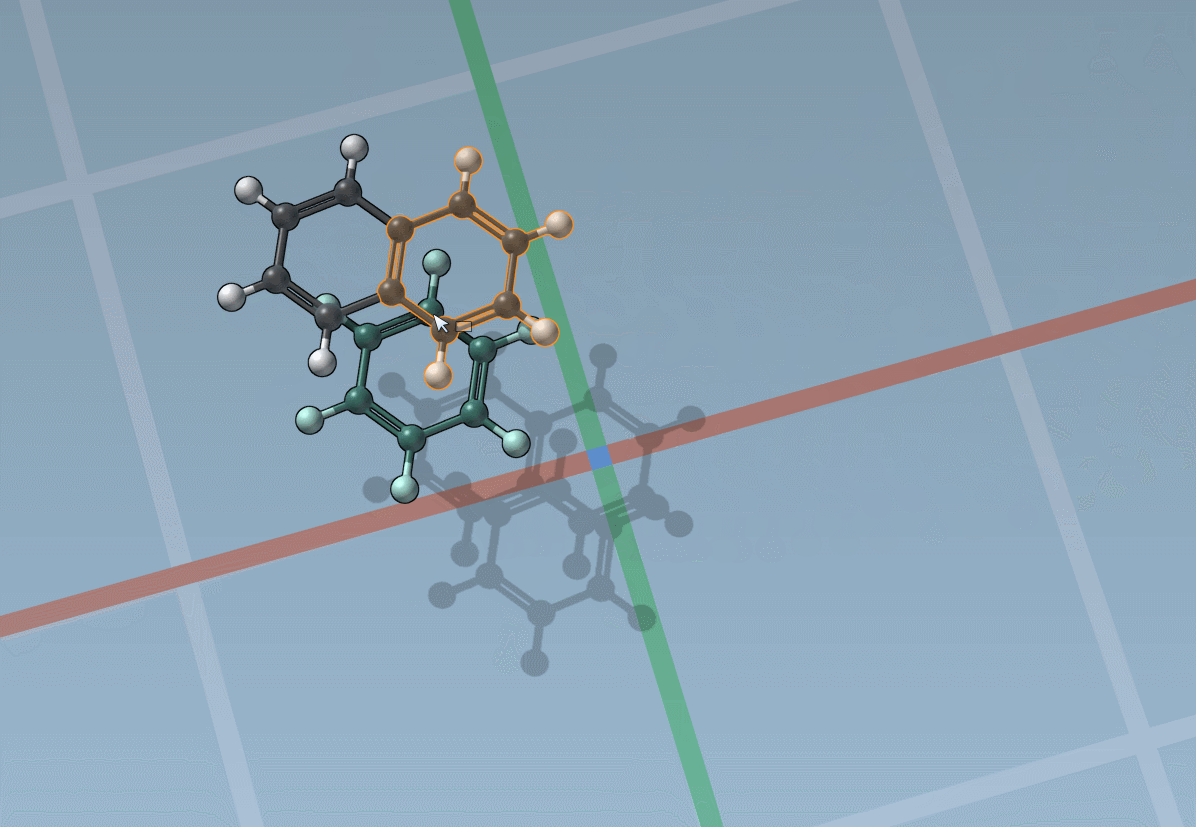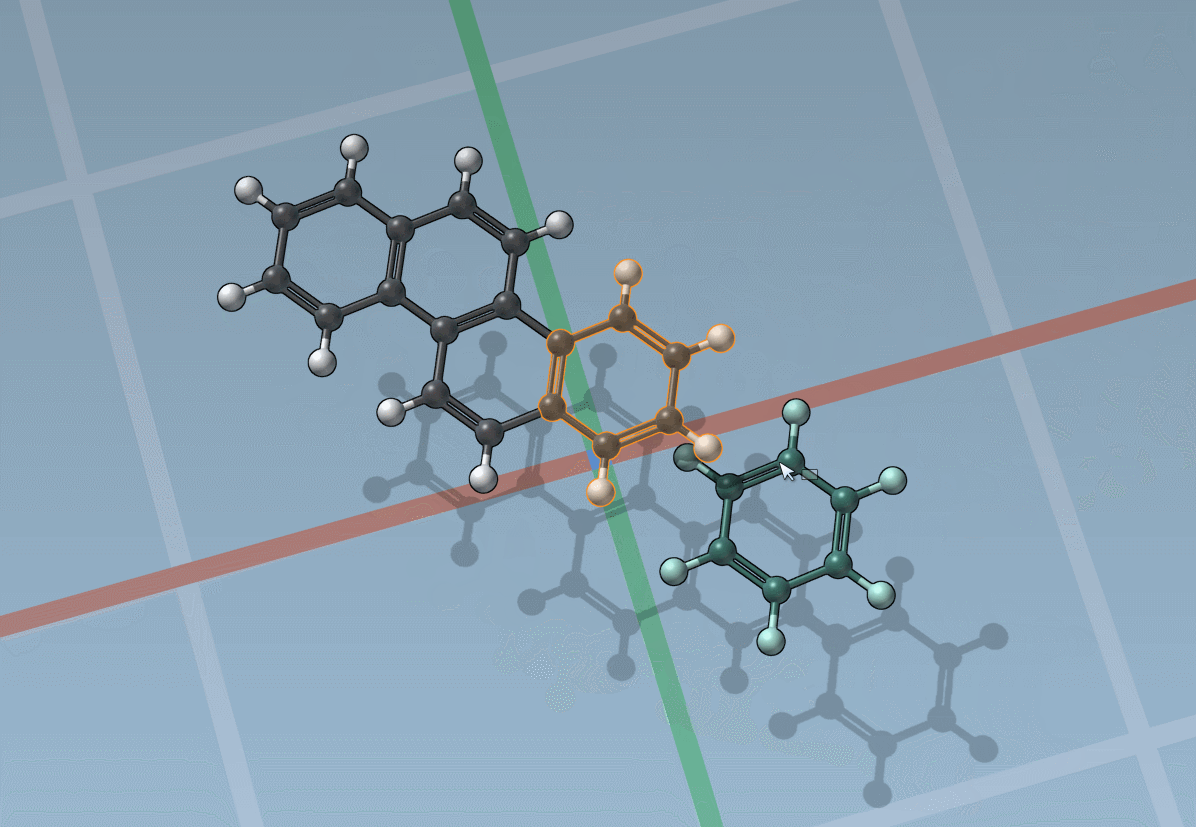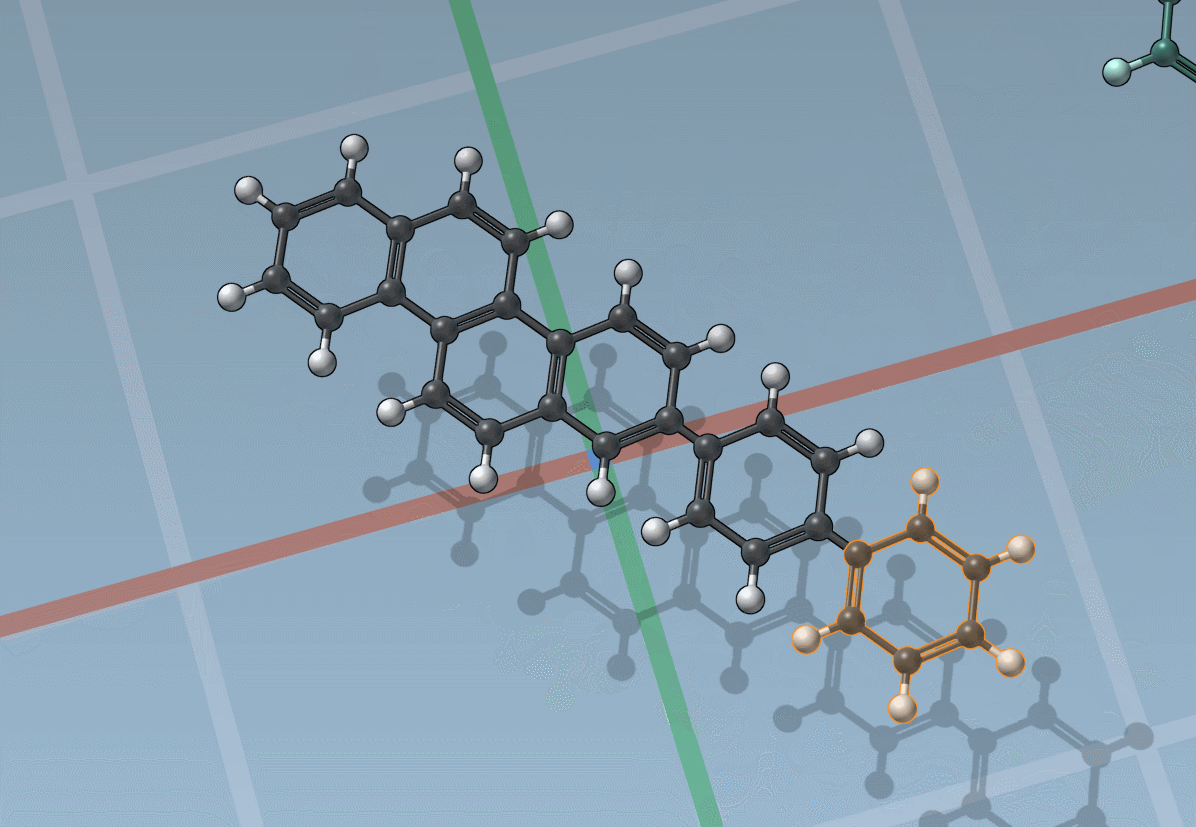
Assets can be anything: rings, fragments, radicals, whole molecules, proteins, nanoparticles, 2D materials, etc.
Just assemble them in the document to rapidly set up complex models for analysis and simulation.
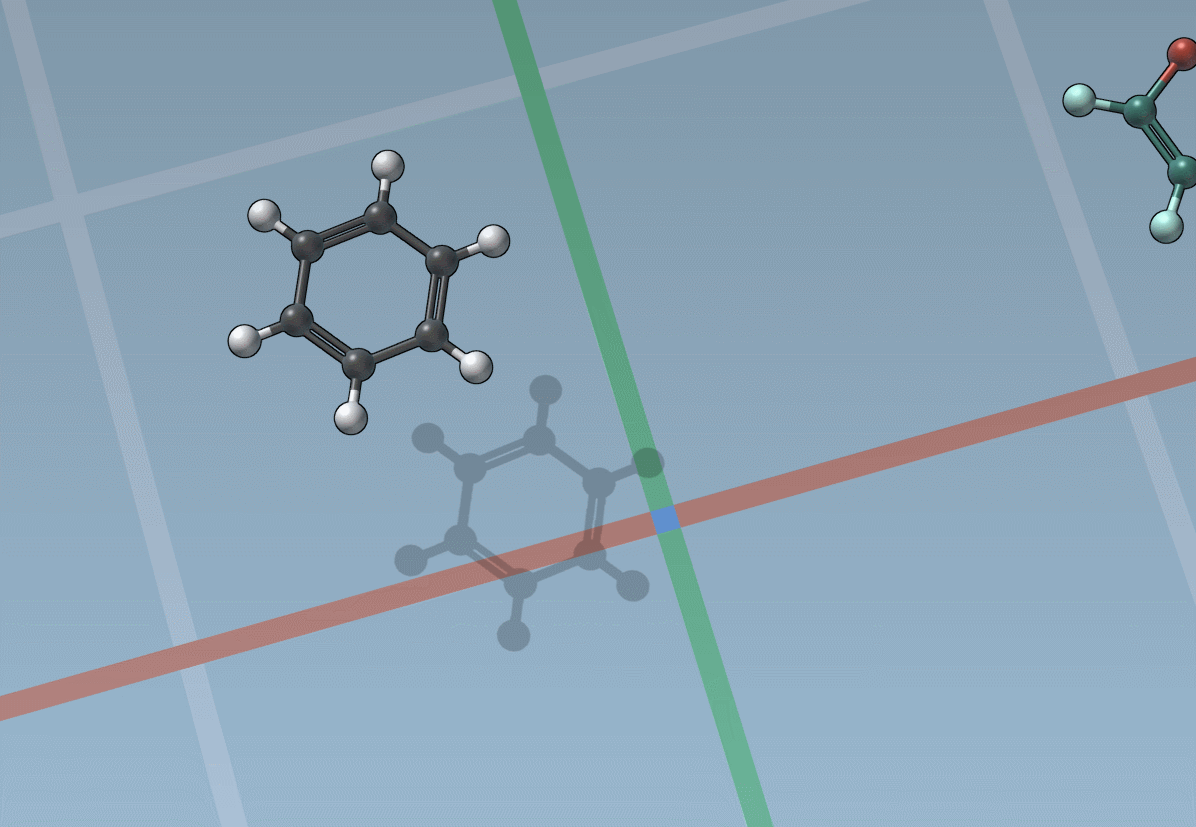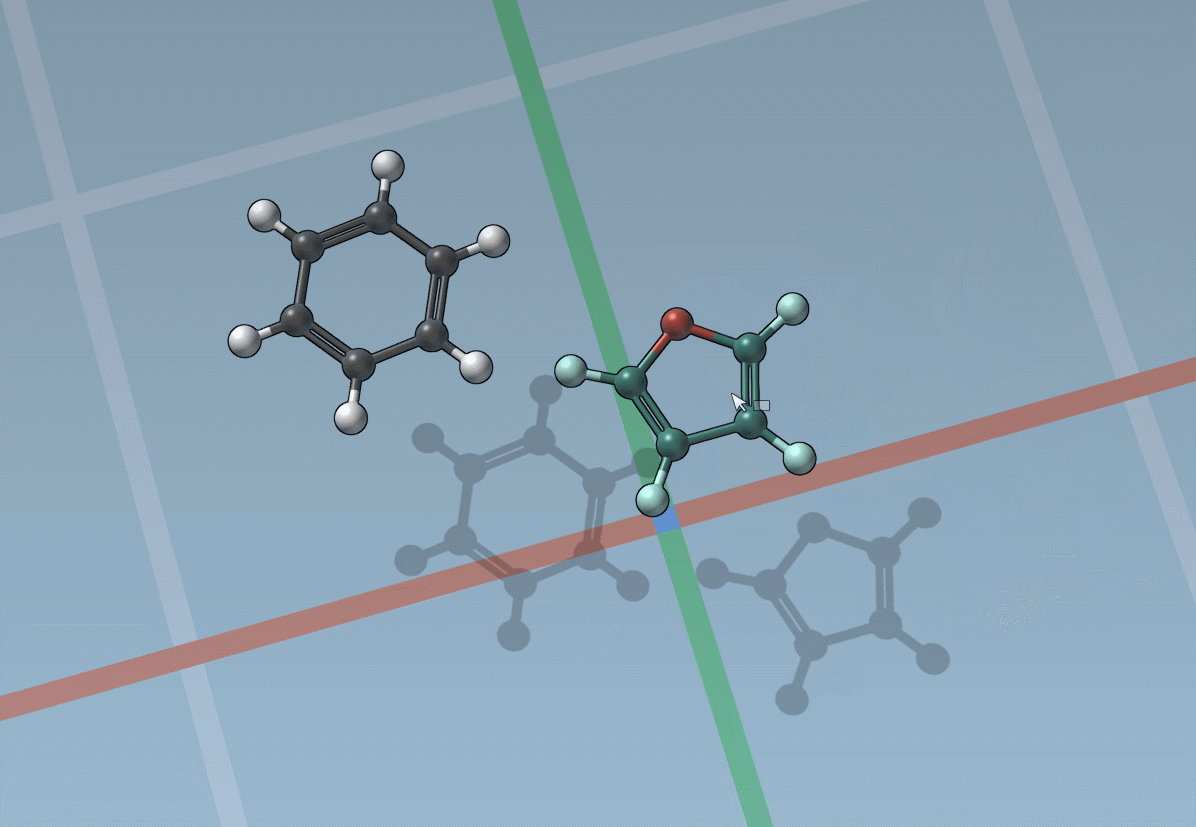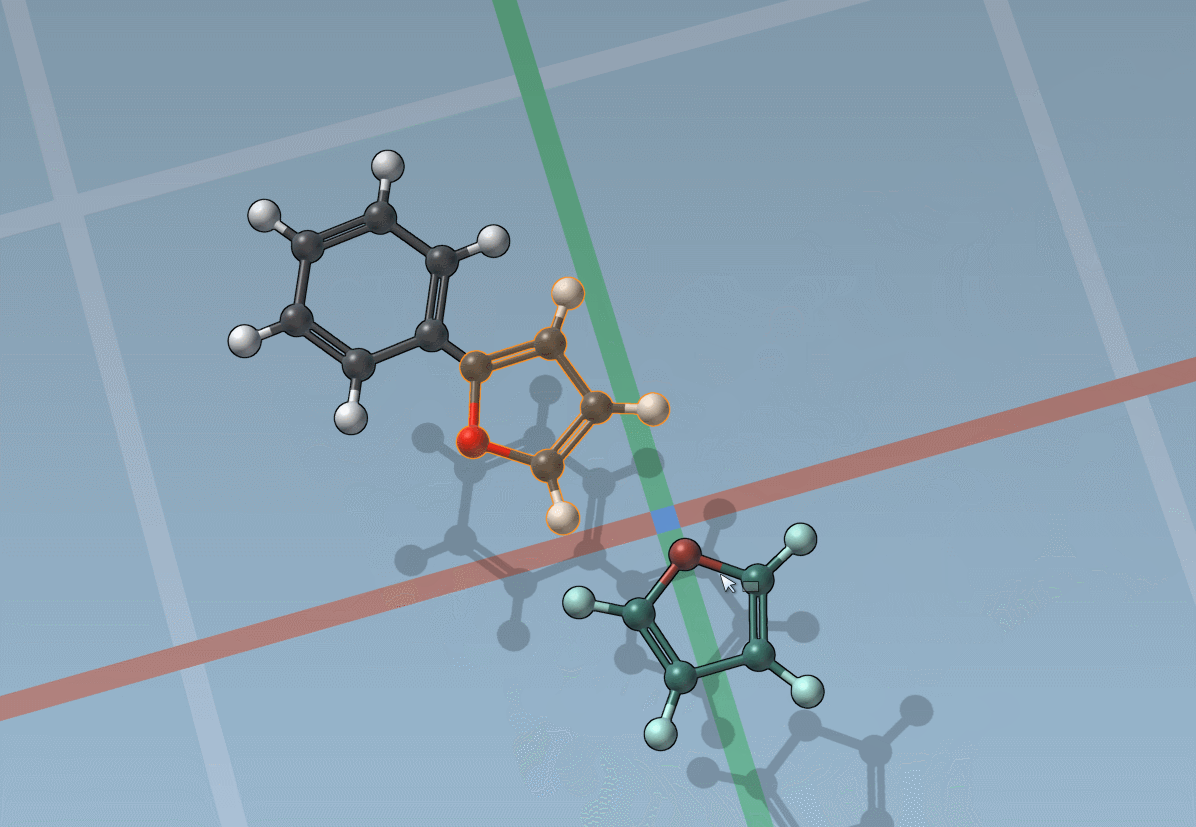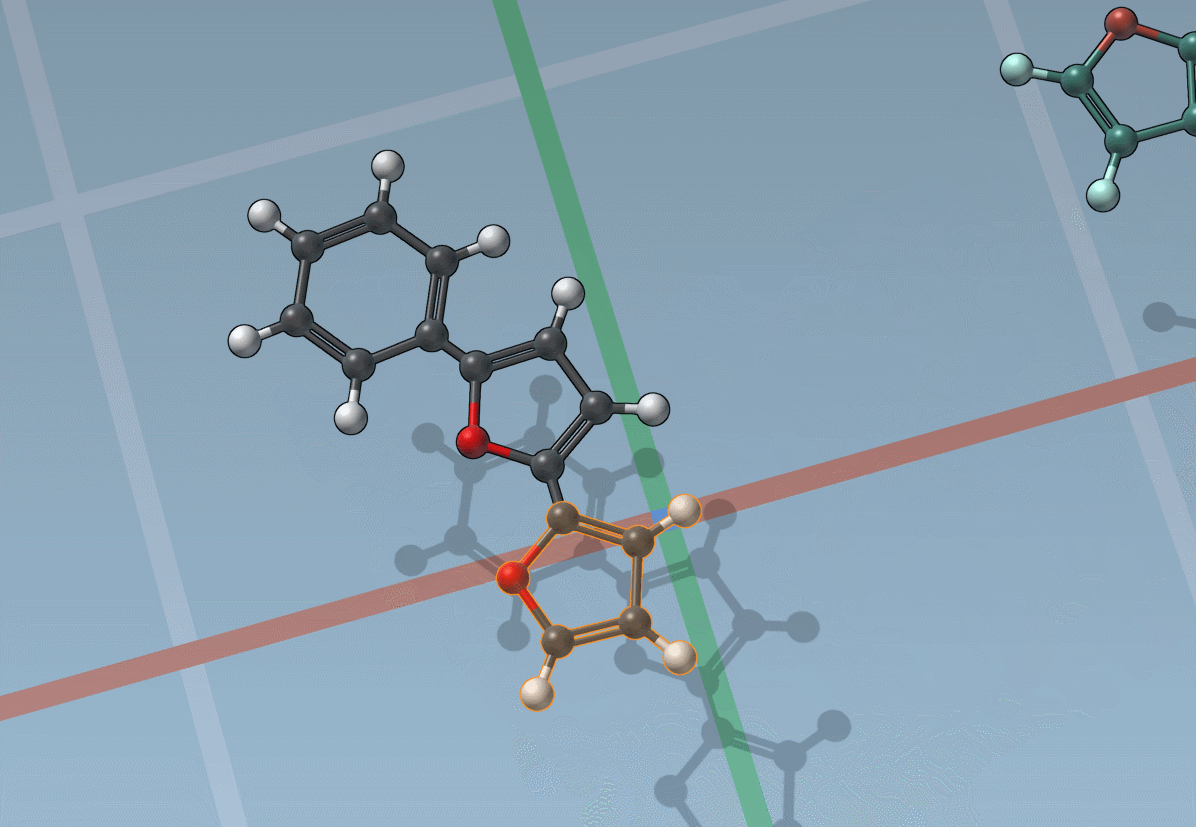
The builder has several cool features to assist you: as shown above, in particular, it has options to predict fragments orientations and prevent implausible substitutions: acceptable substitutions or additions have a green overlay, while forbidden ones have a red overlay (you control in the Preferences whether you want to follow these recommendations or override them).
Whether you’re constructing with individual atoms or with assets, the builder can also adjust hydrogens for you, and merge overlapping atoms.
By pressing and holding the Shift key, you can choose which atom and bond in the asset are used to replace the atoms and bonds you click in the document:
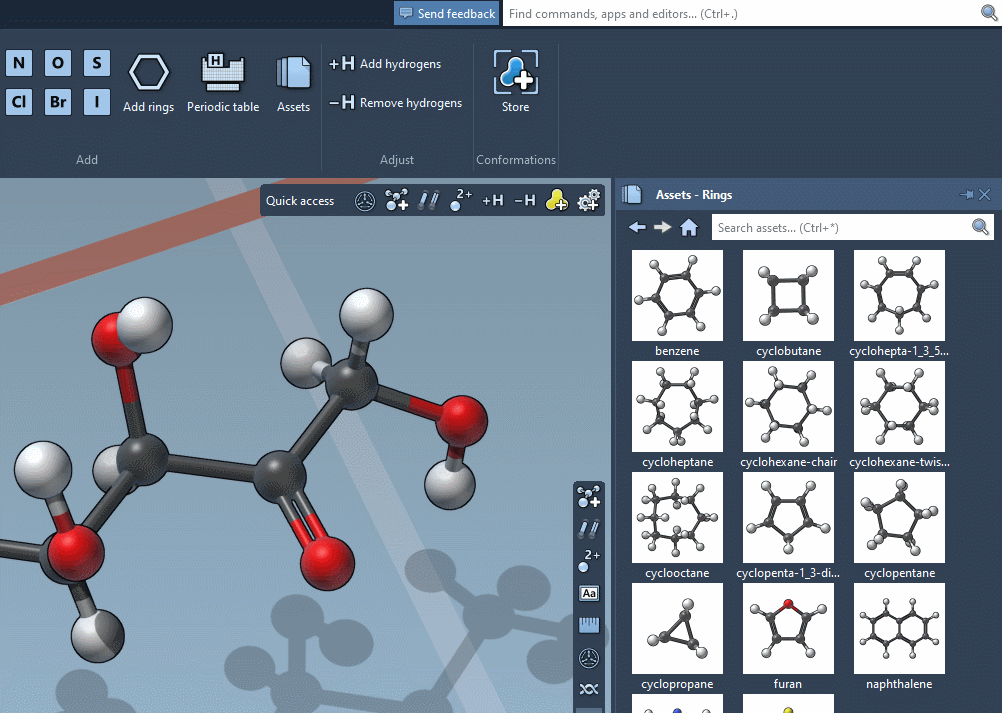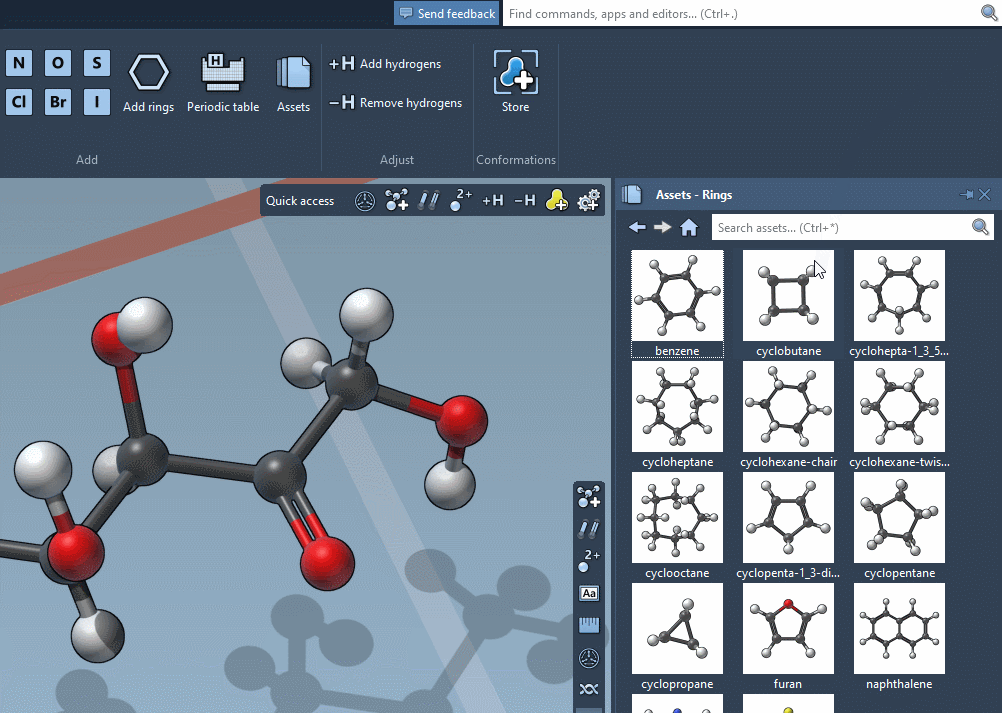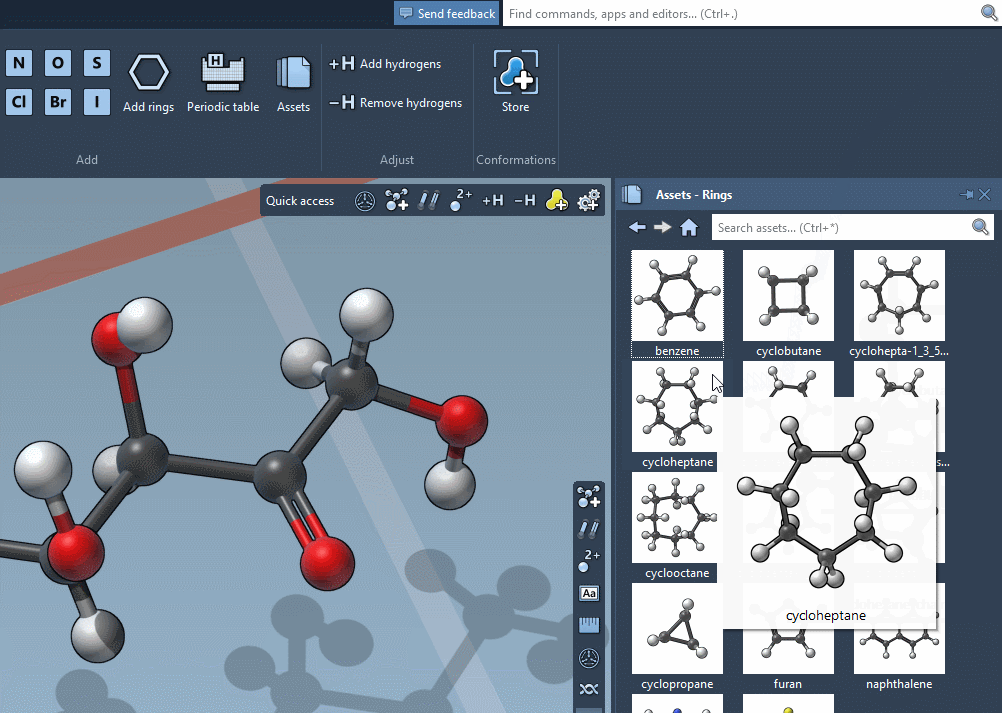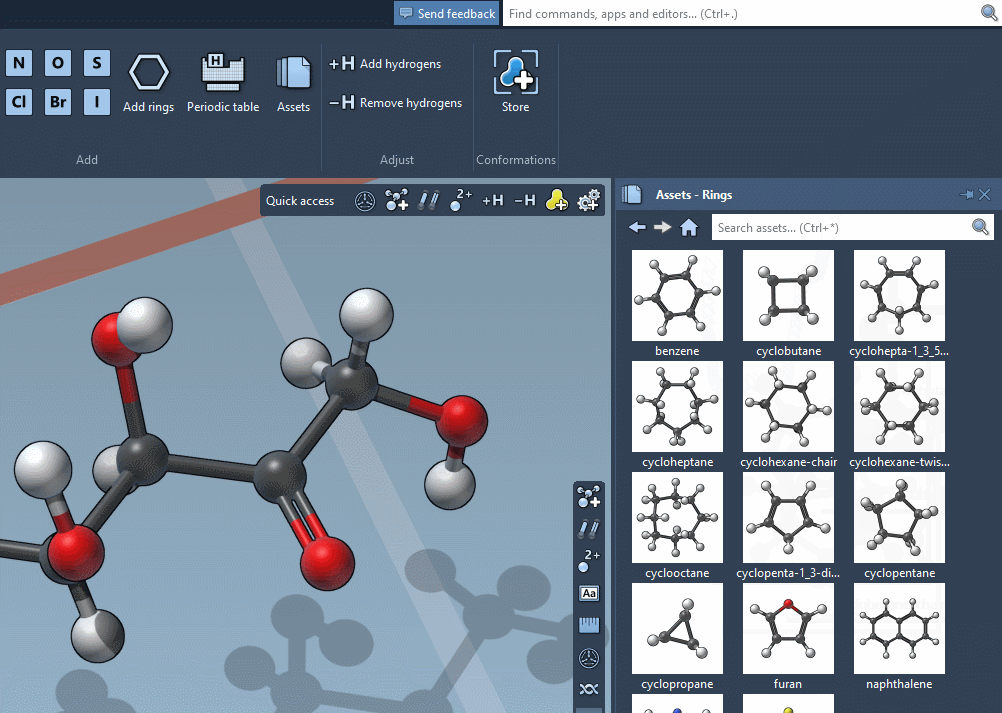
Accordingly, the SAMSON interface has a new component, the Asset Browser, which gathers the assets included by default in SAMSON with the assets you obtain from SAMSON Connect:
B is for “Bonds”
Of course, we made it easy to edit bond orders:
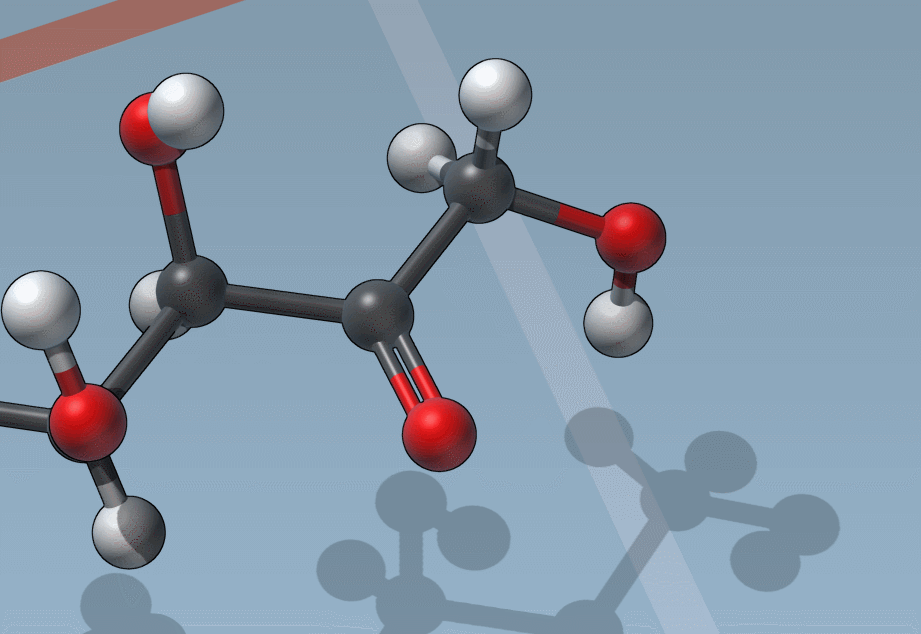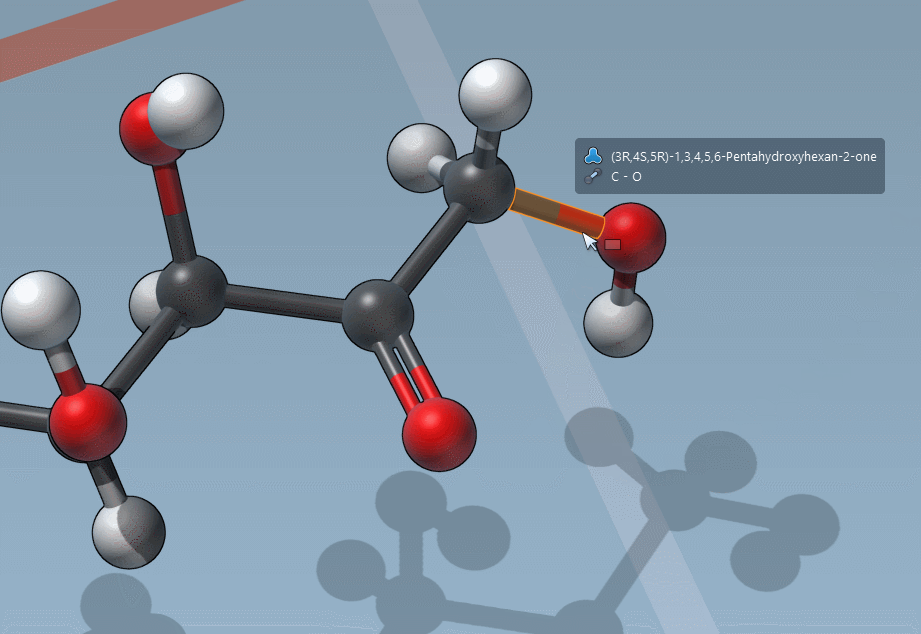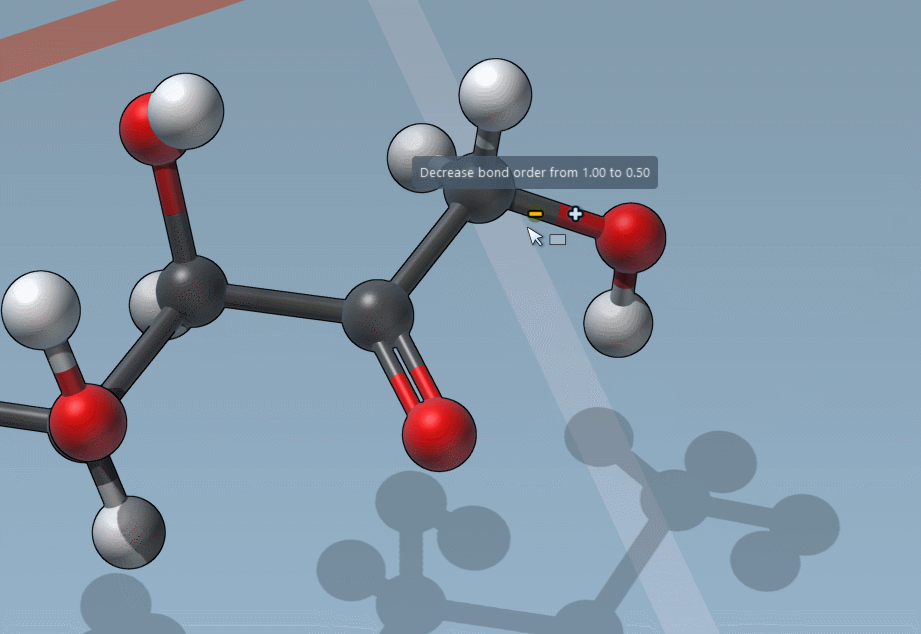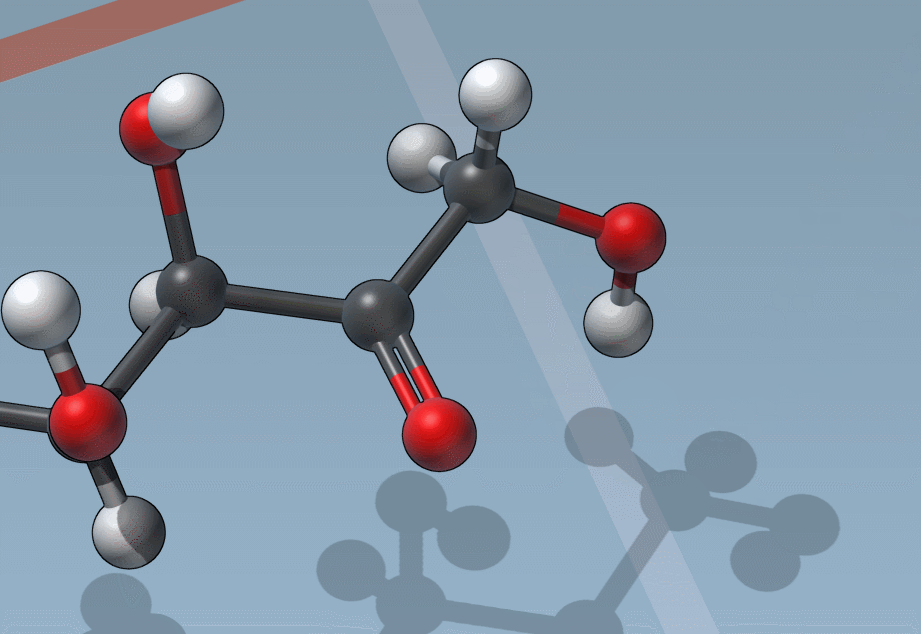
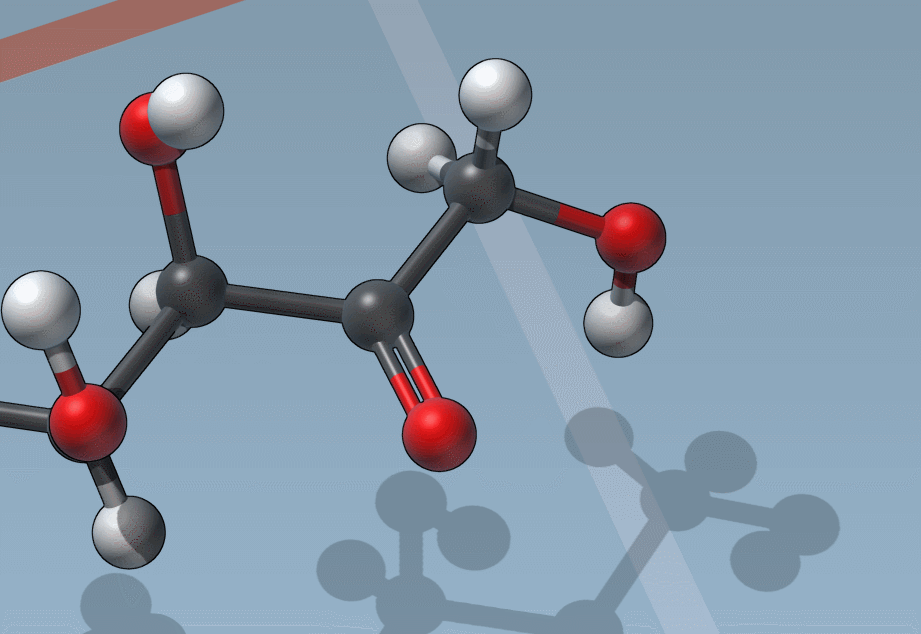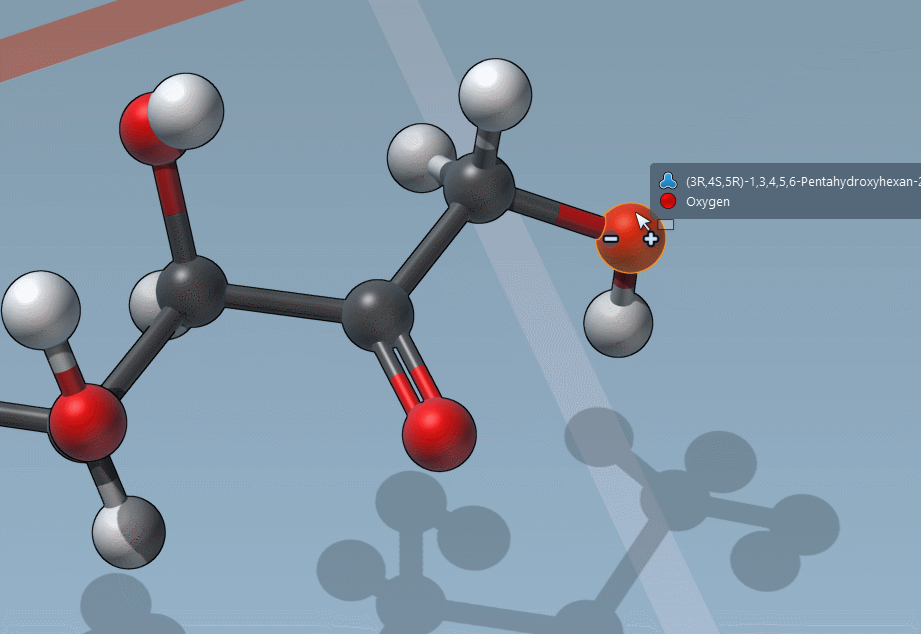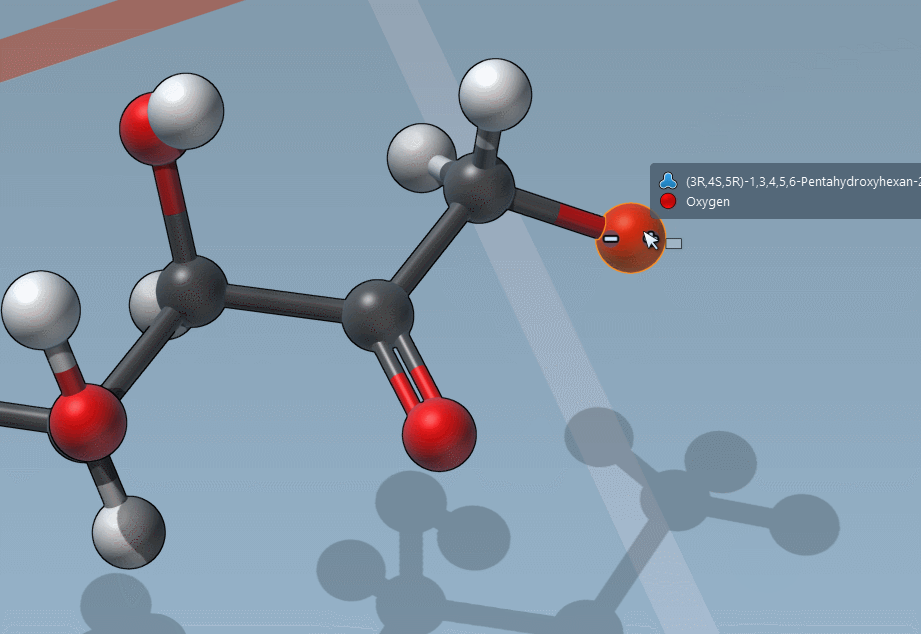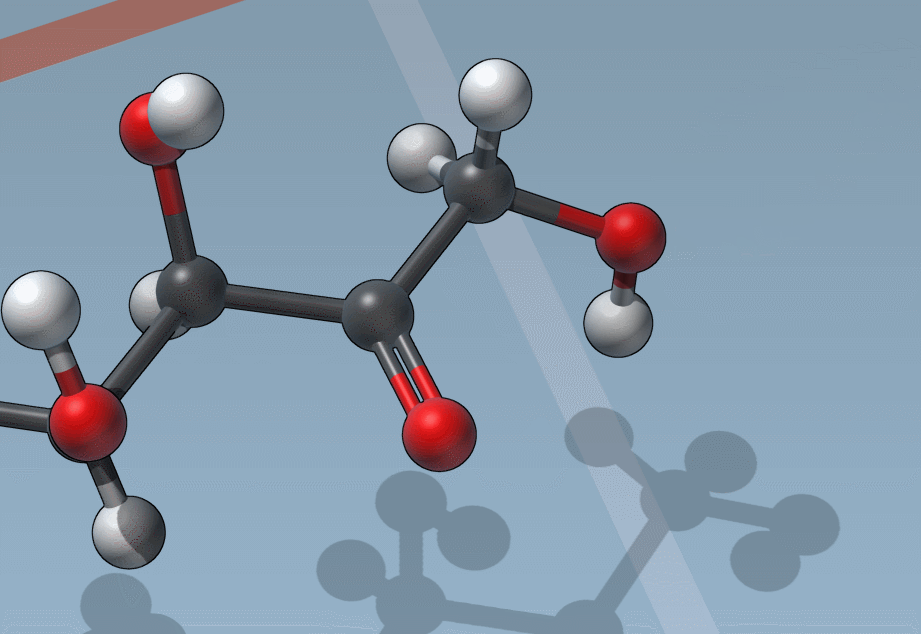
C is for “Charges”
And it’s as easy to change formal charges:
You like to move it? Move it!
You now have precise control over positions and orientations of molecules using the Move editor:
You can also control the pivot of the Move editor:
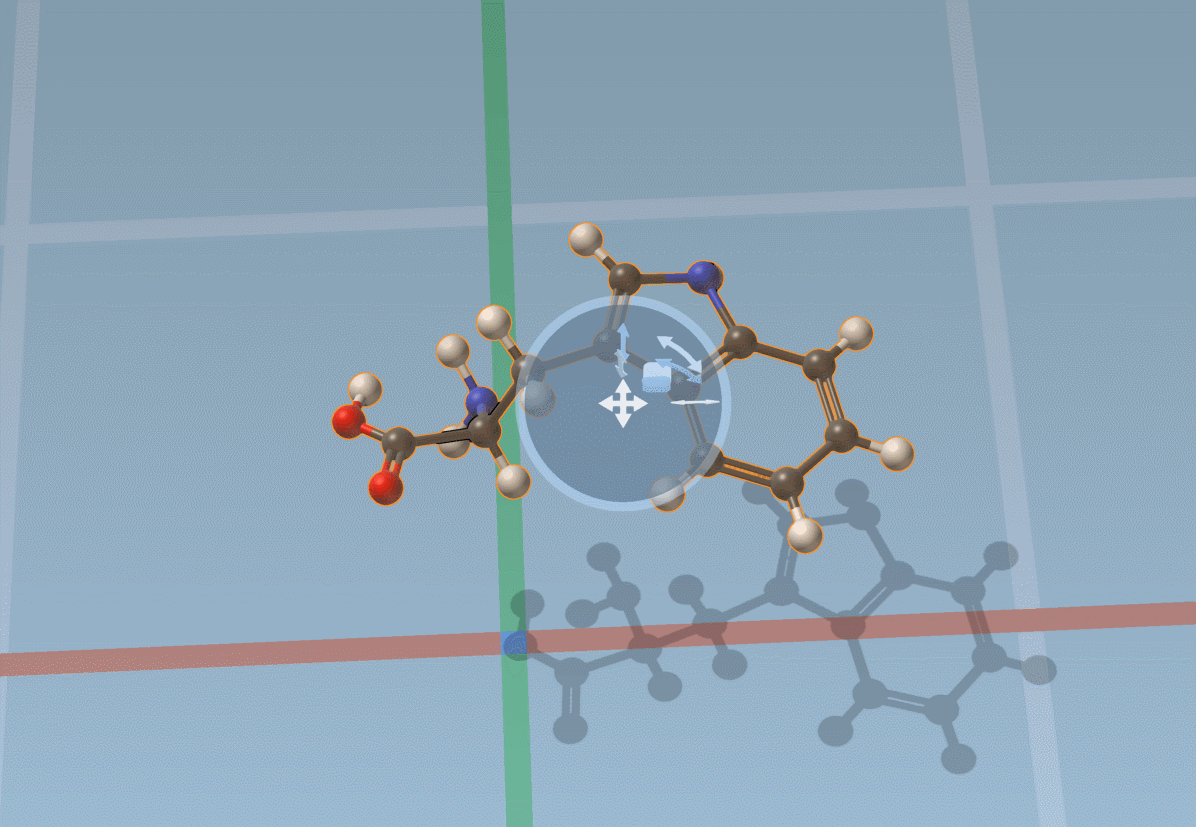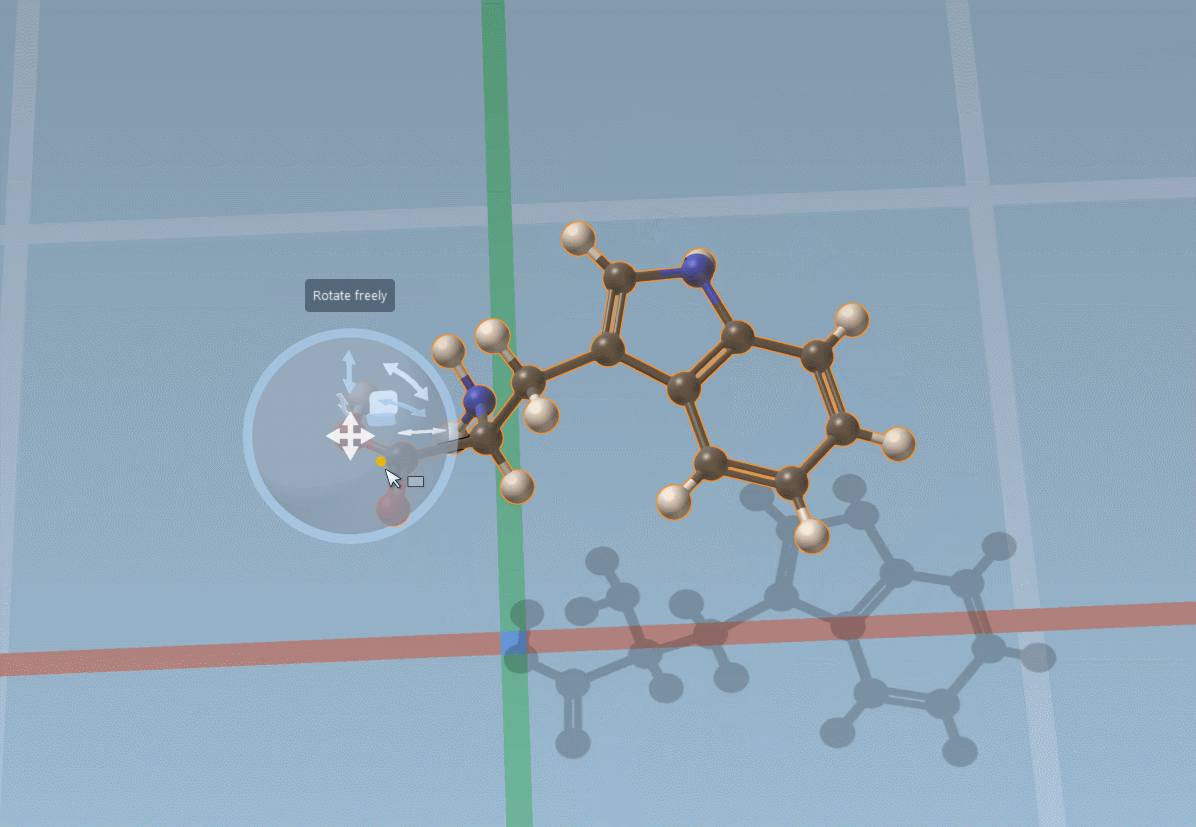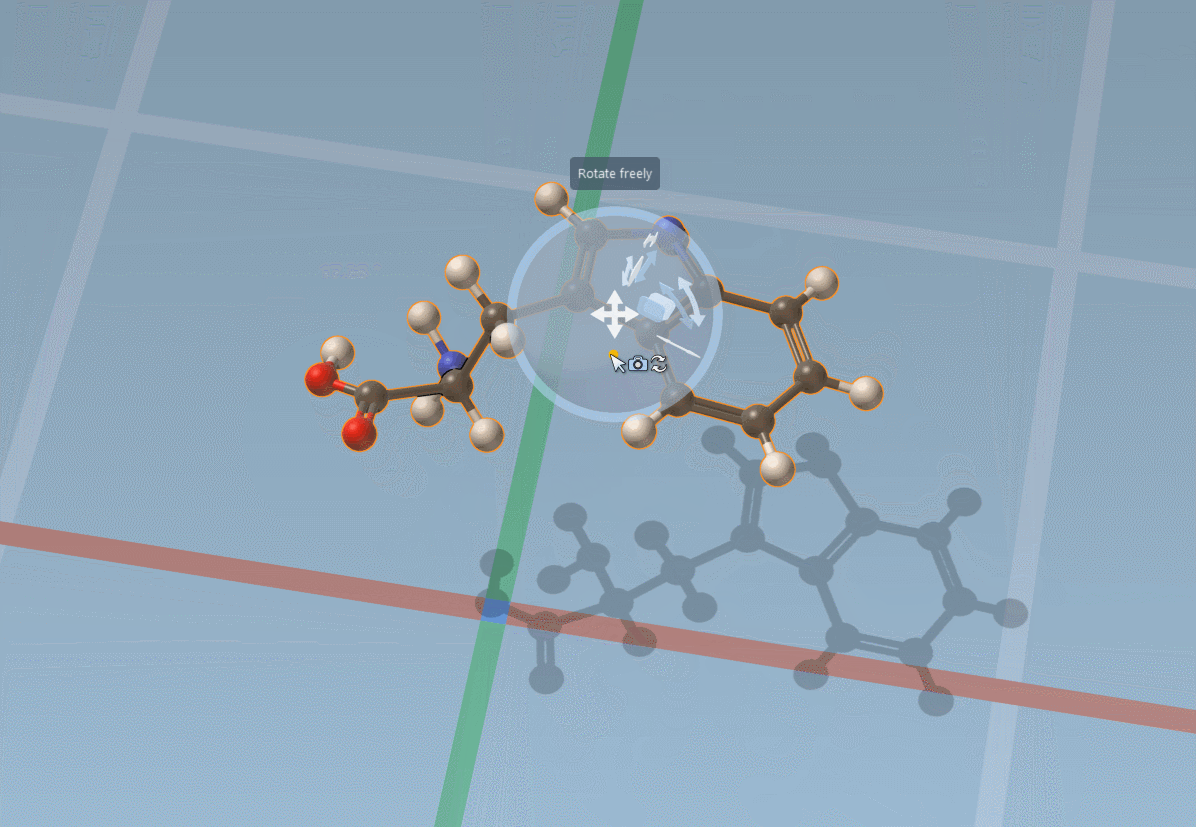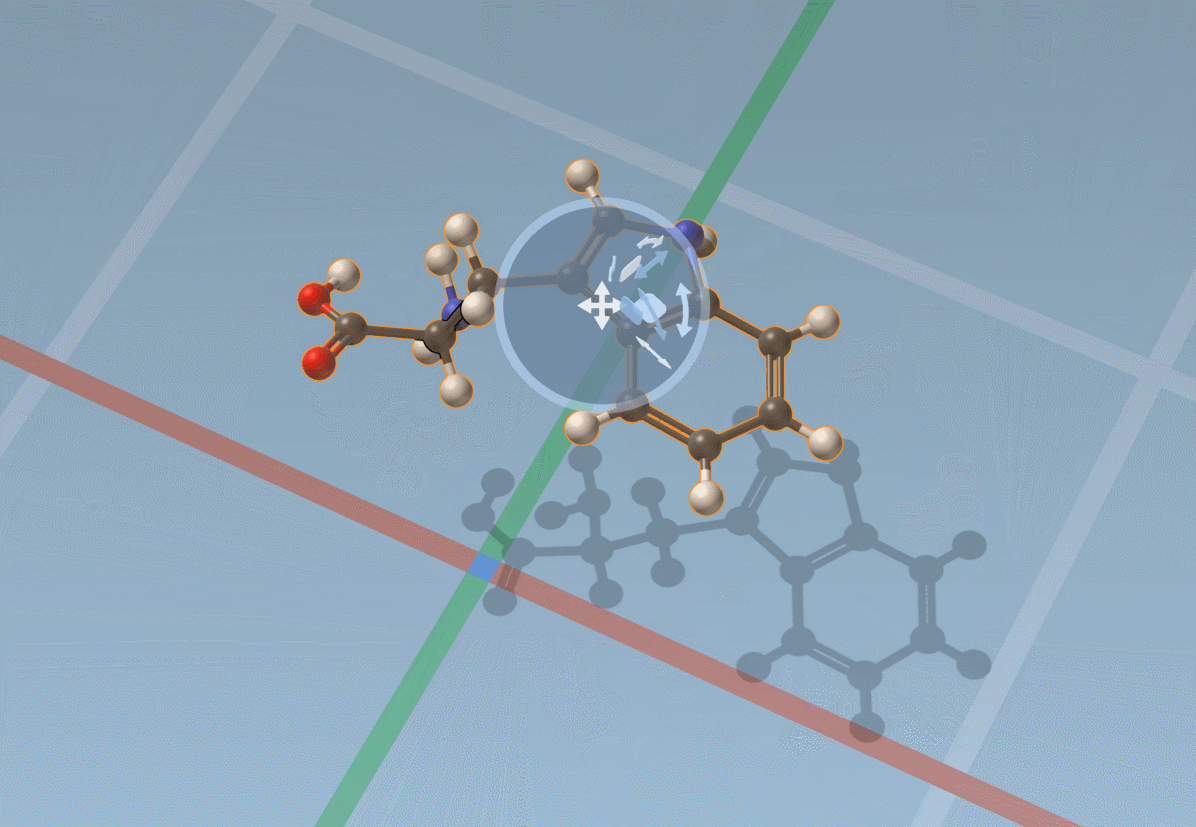
The Move editor can also be used to position parts of molecules. In this case, it does its best to automatically find the best pivot and orientation:
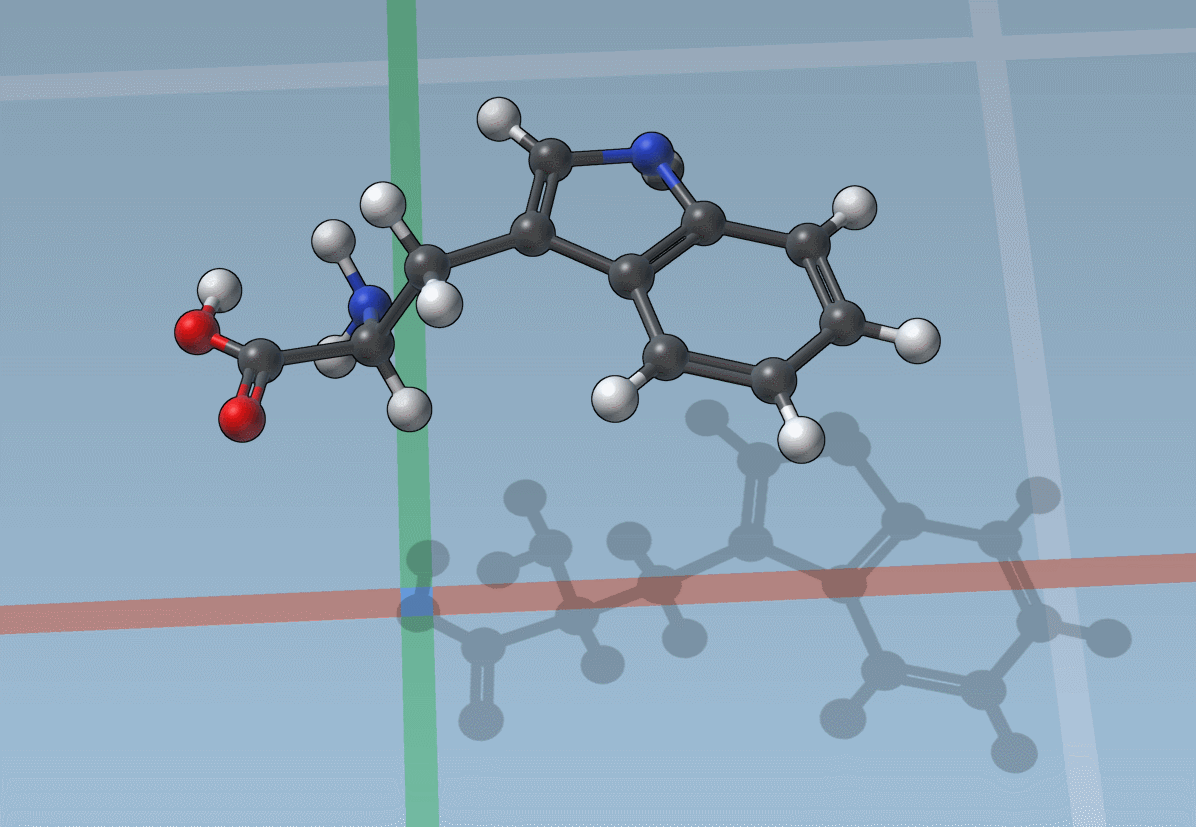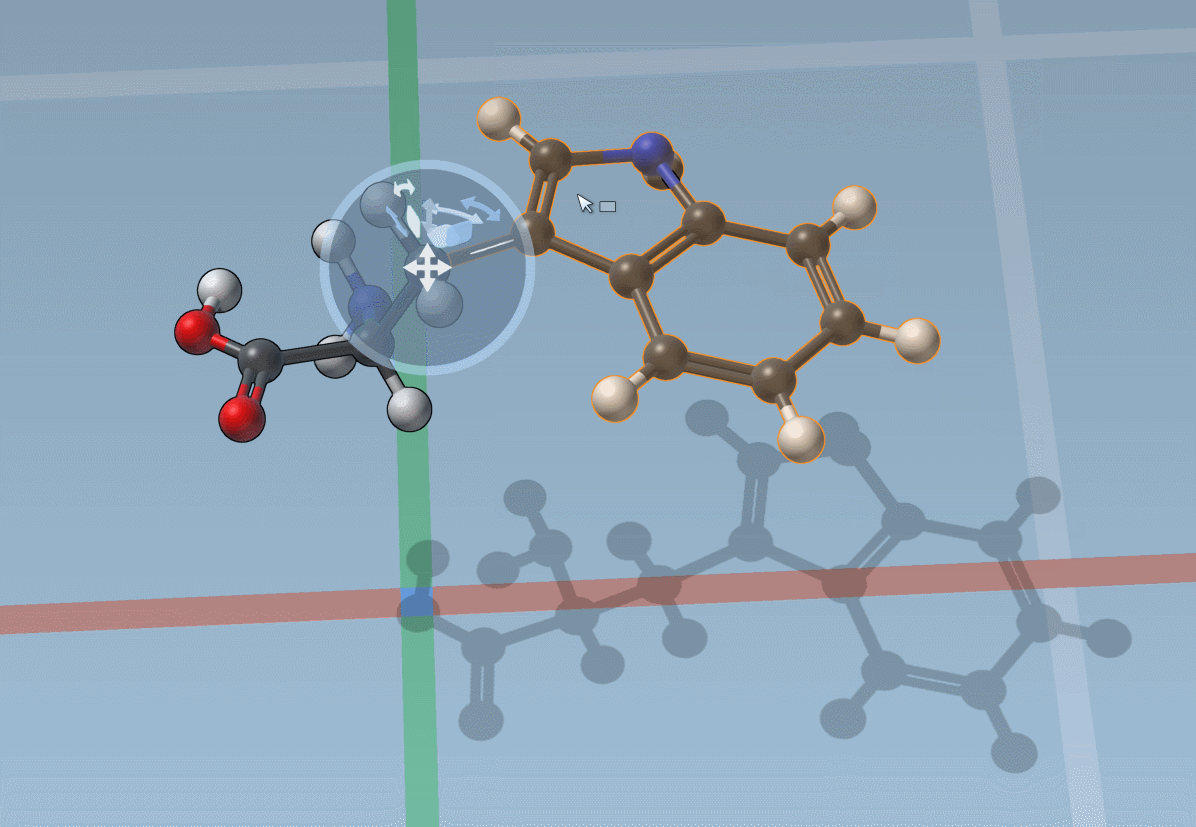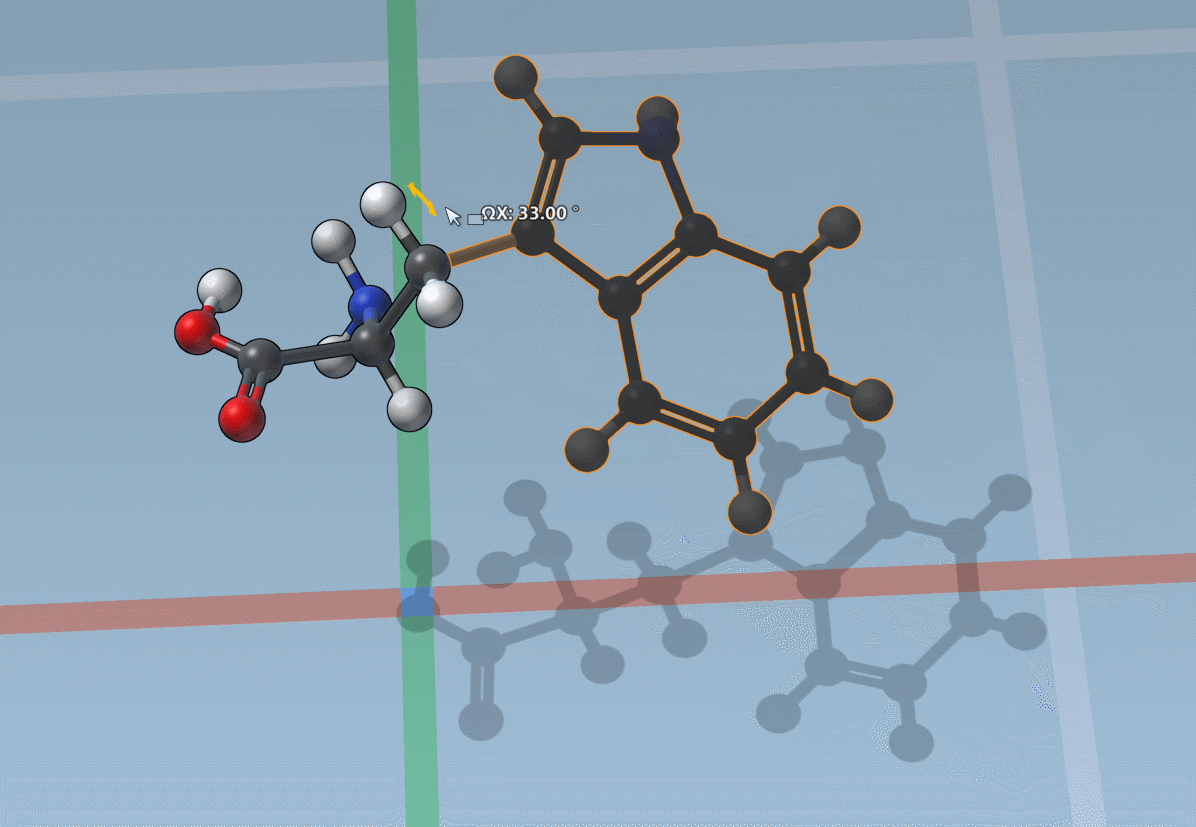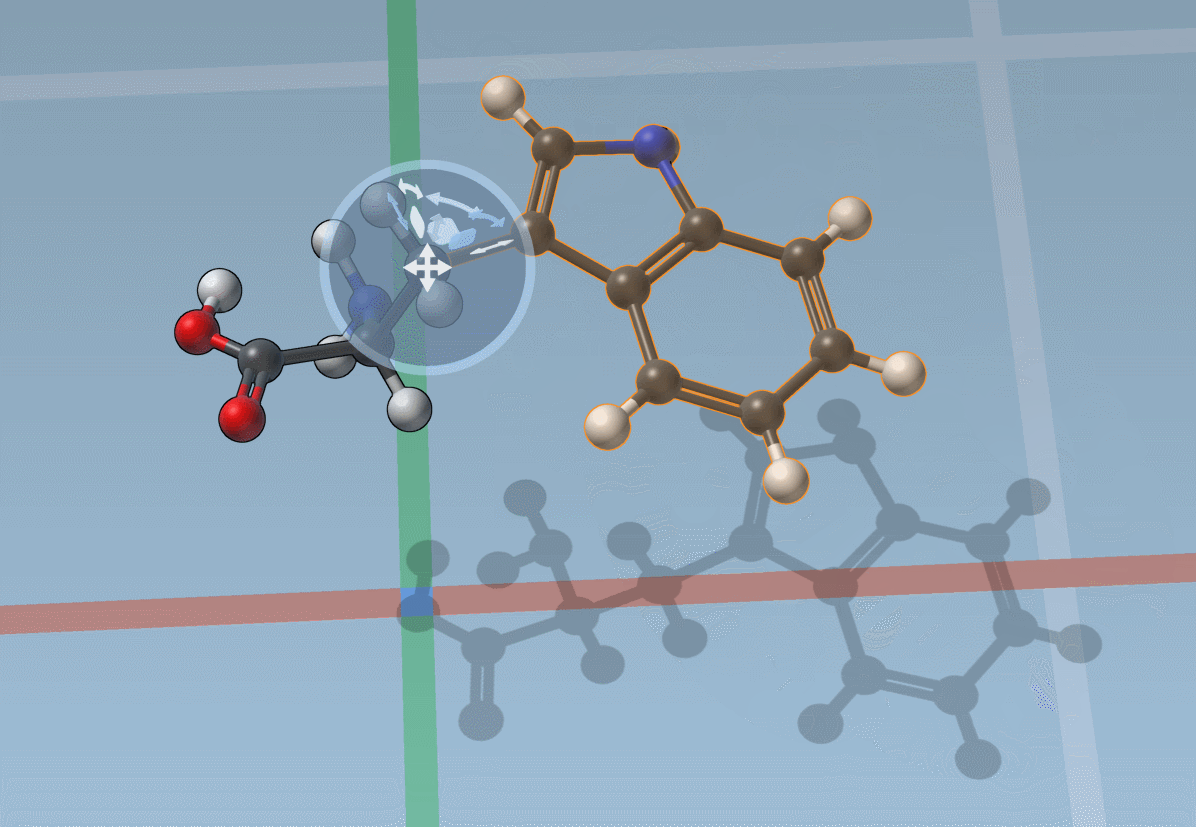
You can also snap translations and rotations to precise values:
Of course, you can use the Move editor when performing interactive energy minimization and simulation:
And if you want to check how much things have moved, the Measure editor has been completely redesigned and makes it easy to measure distances, angles and dihedrals:
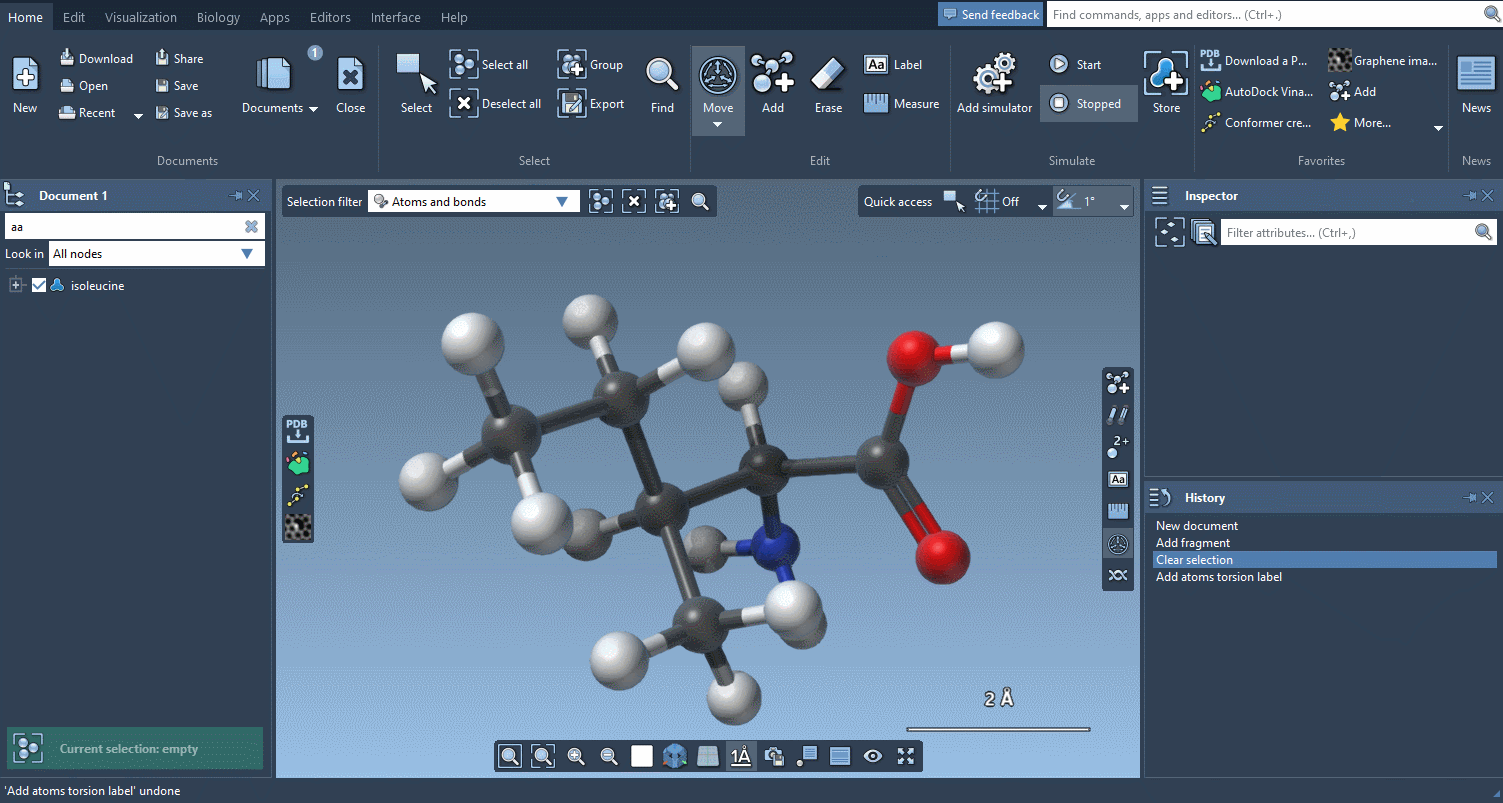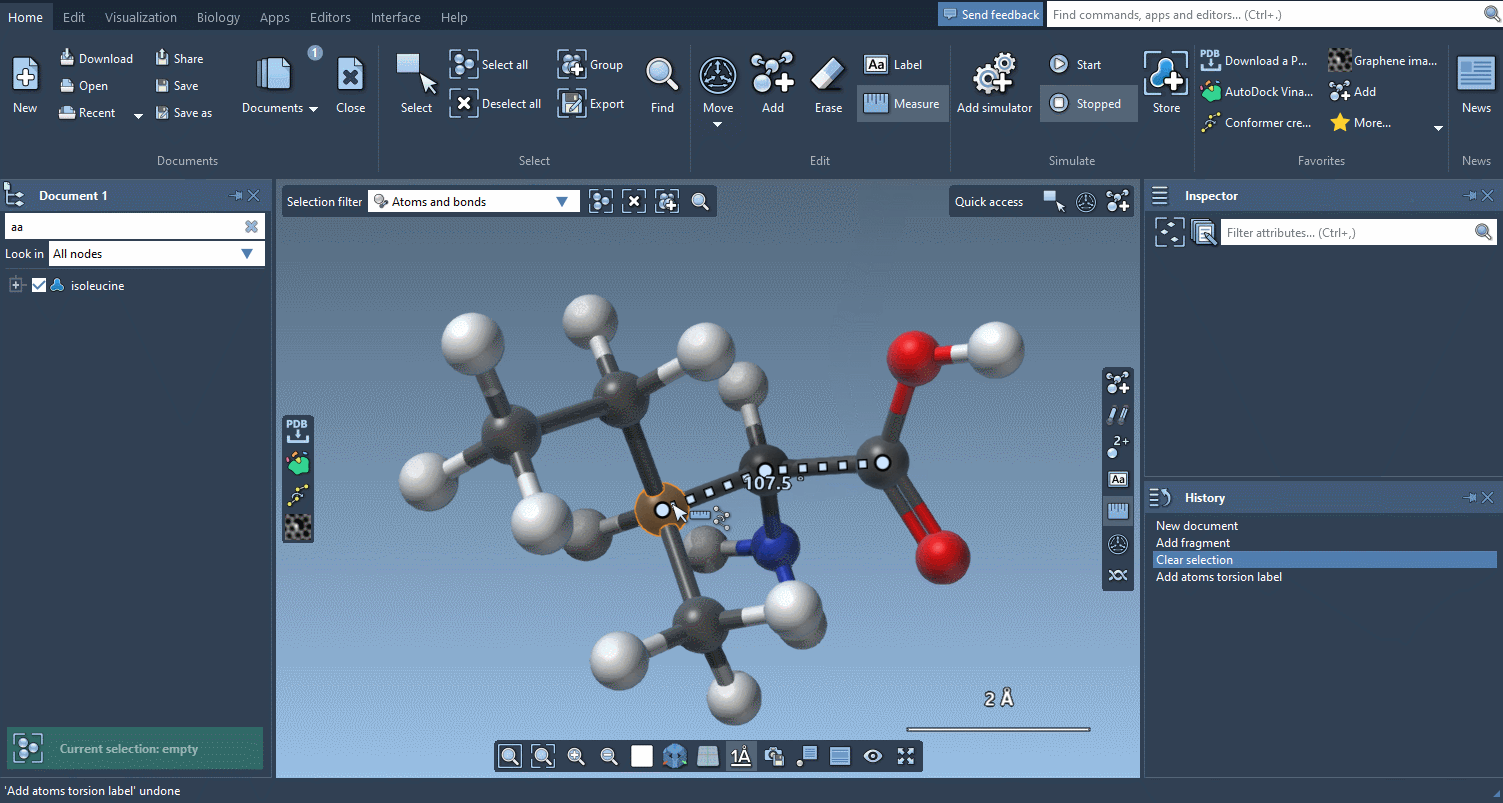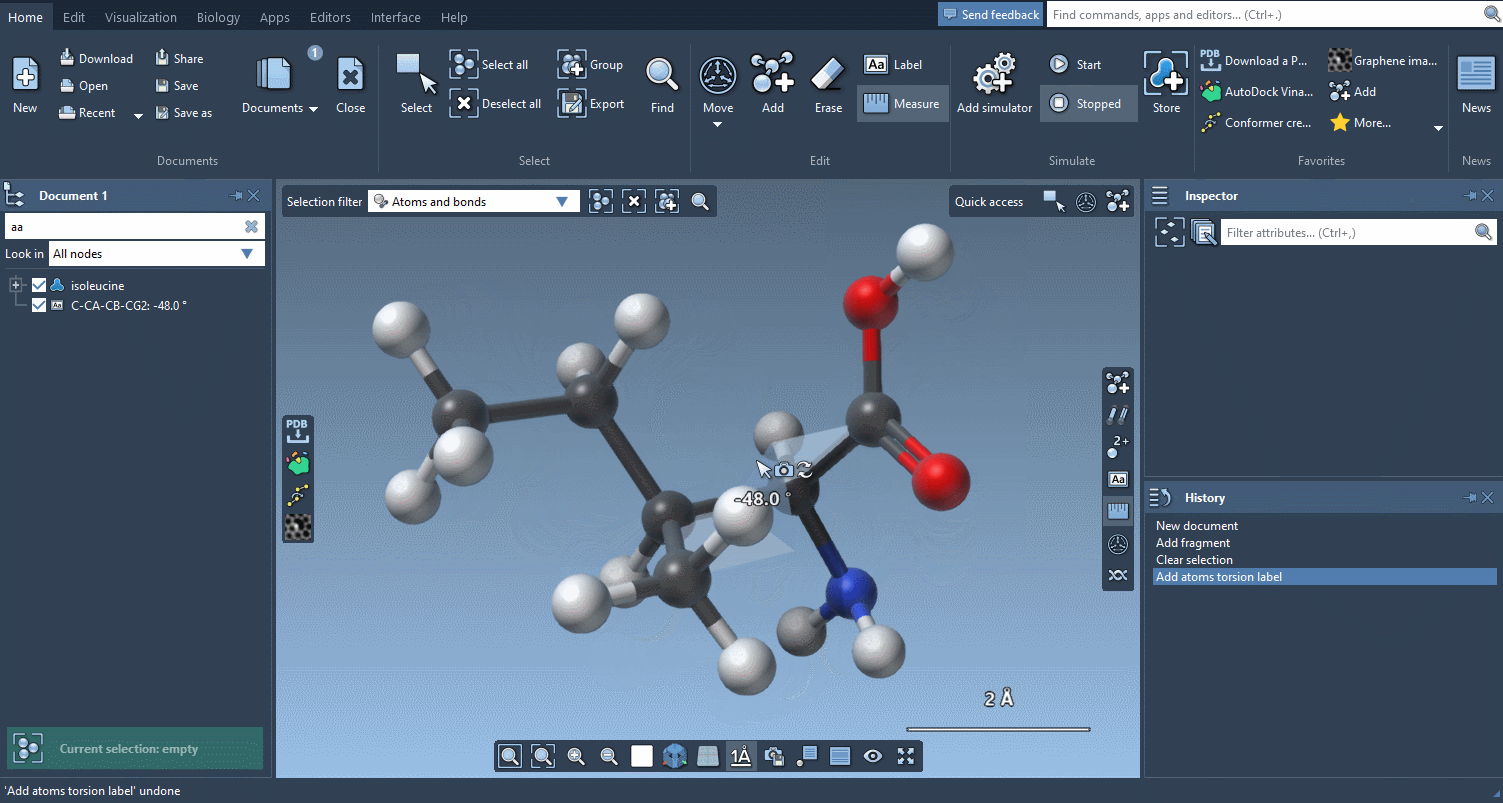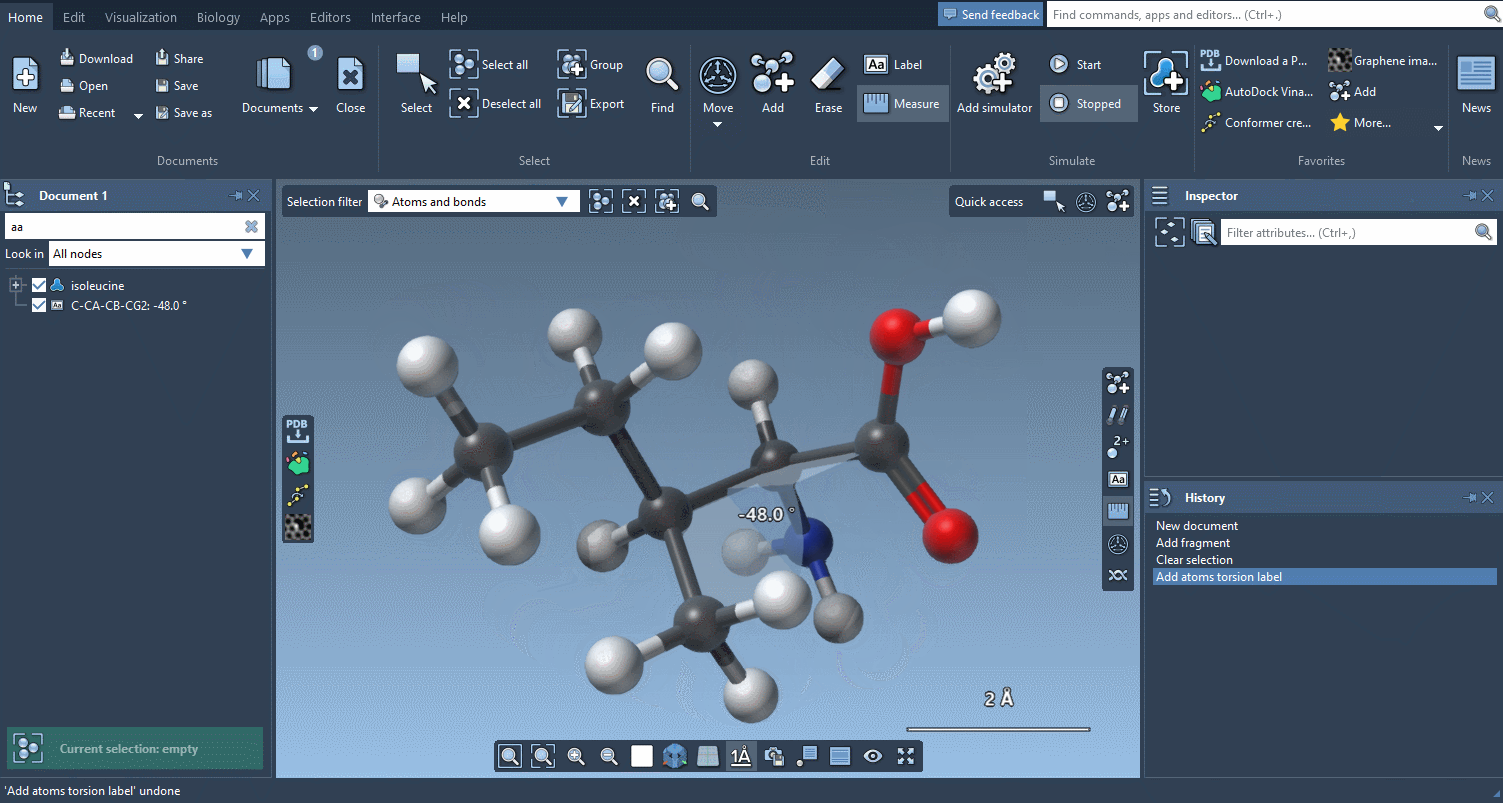
An improved user experience
Following user feedback, the menu was significantly updated to make frequent tasks easily accessible:
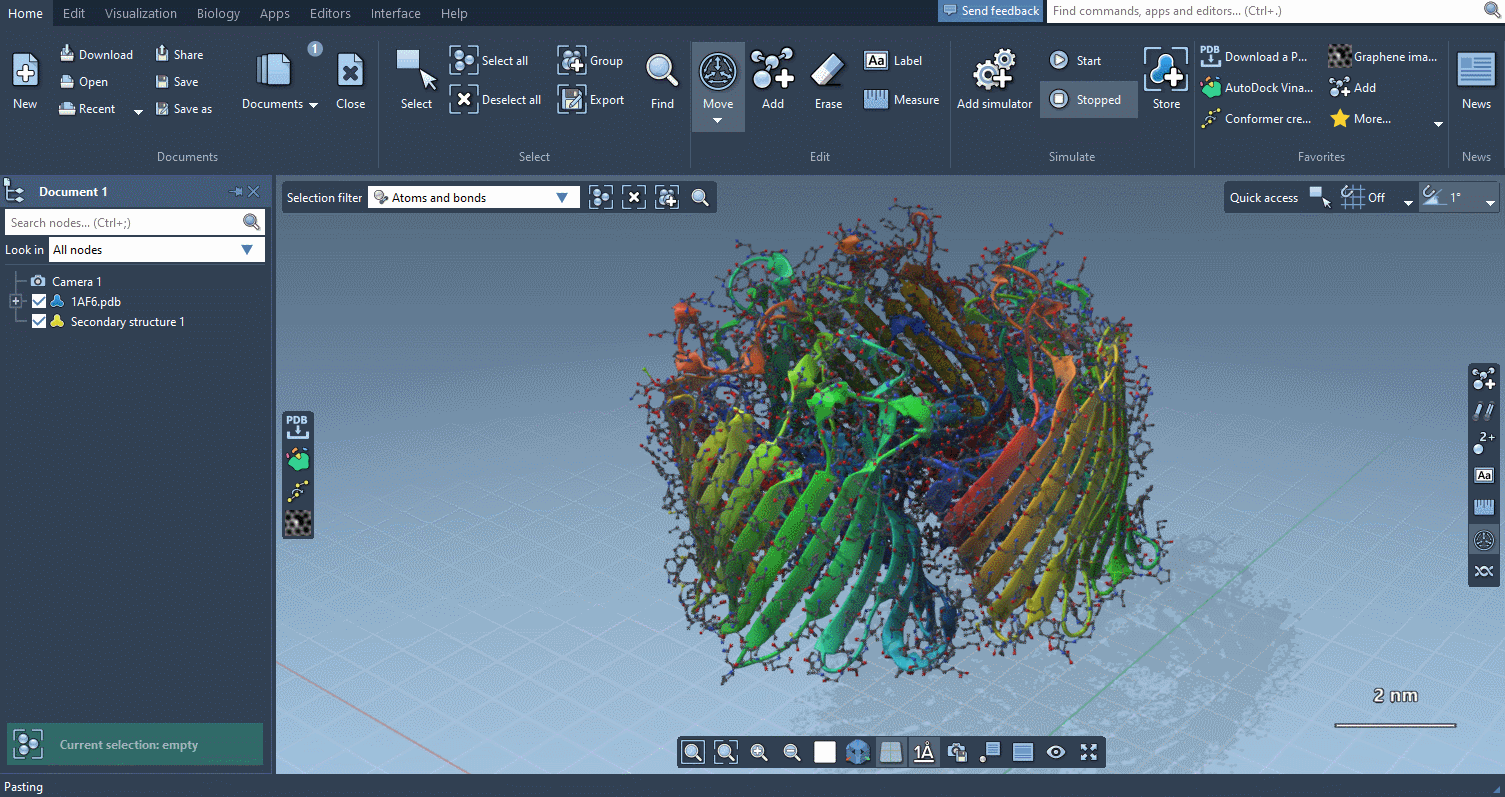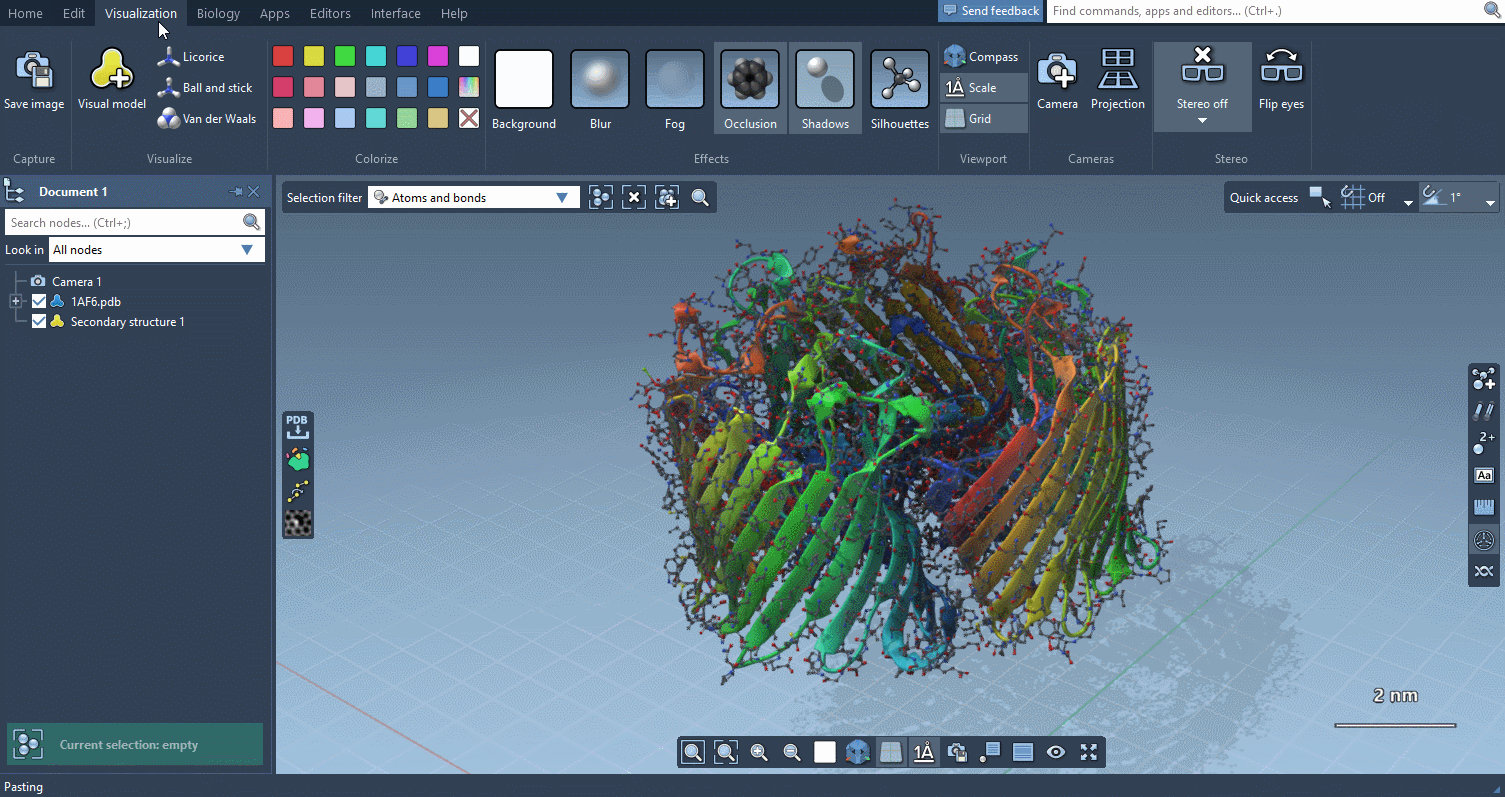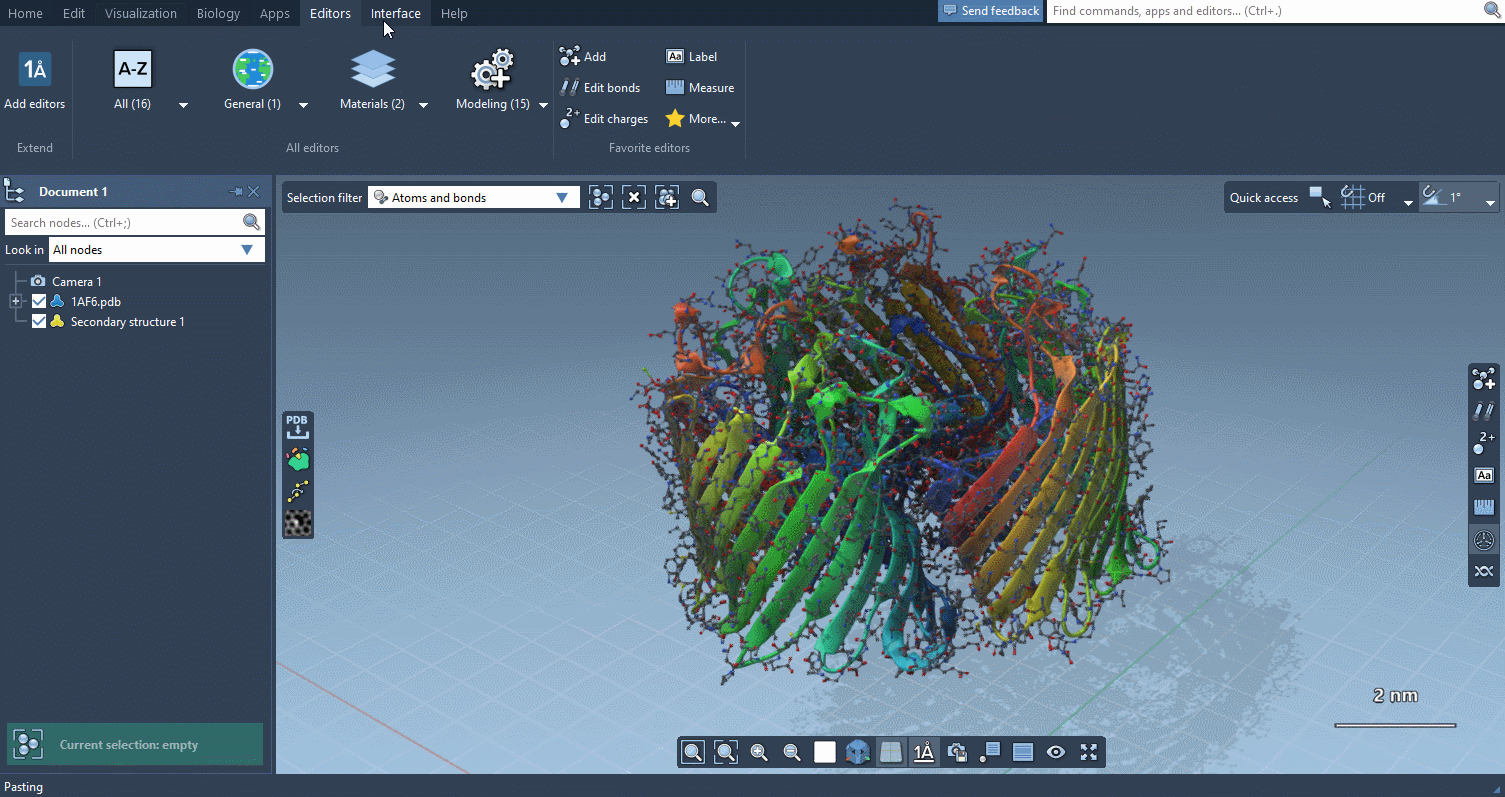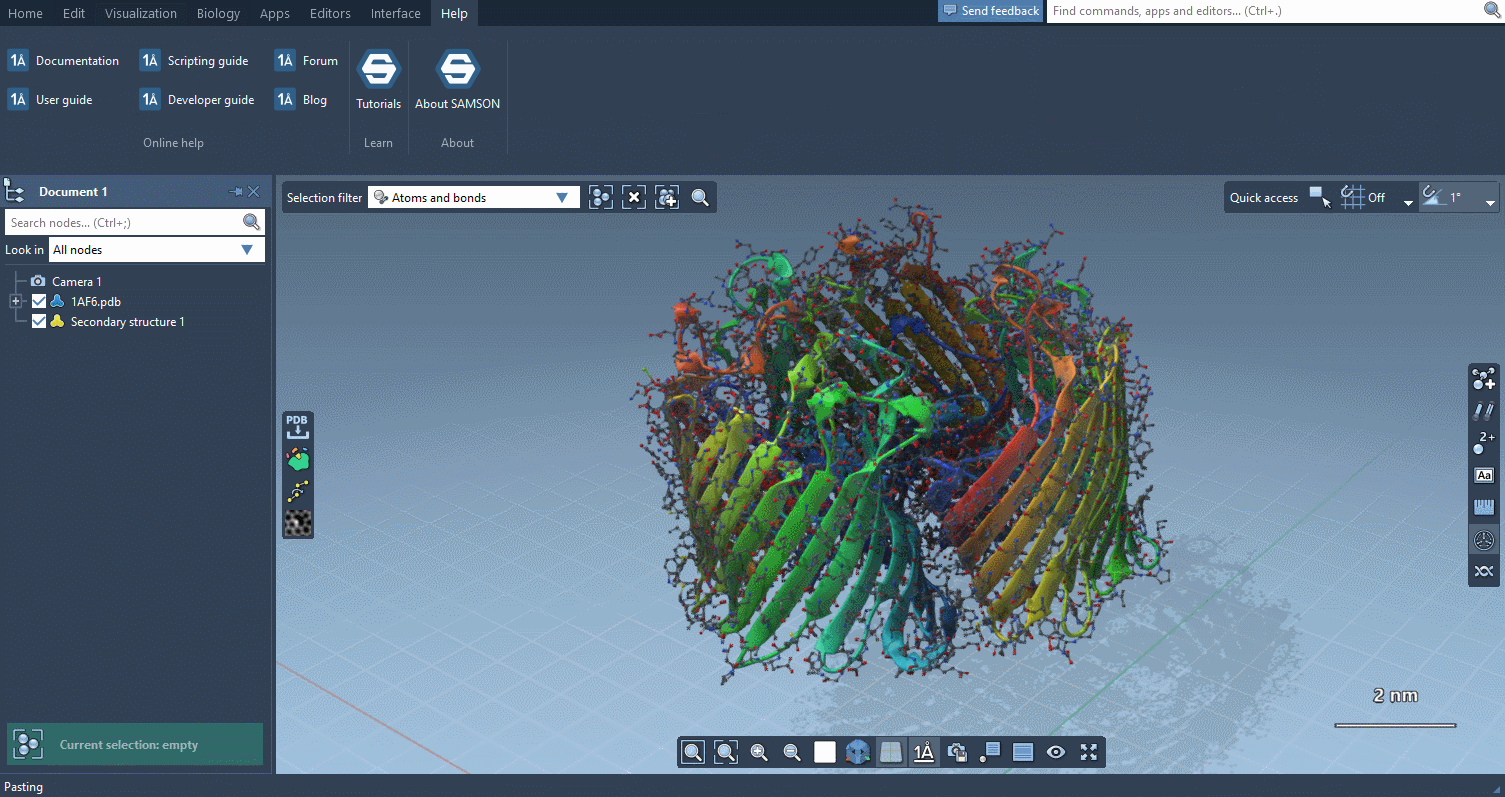
Visualization menu
The Visualization menu now makes it very easy to apply traditional molecular representations, change colors, and control advanced rendering options with a few clicks:
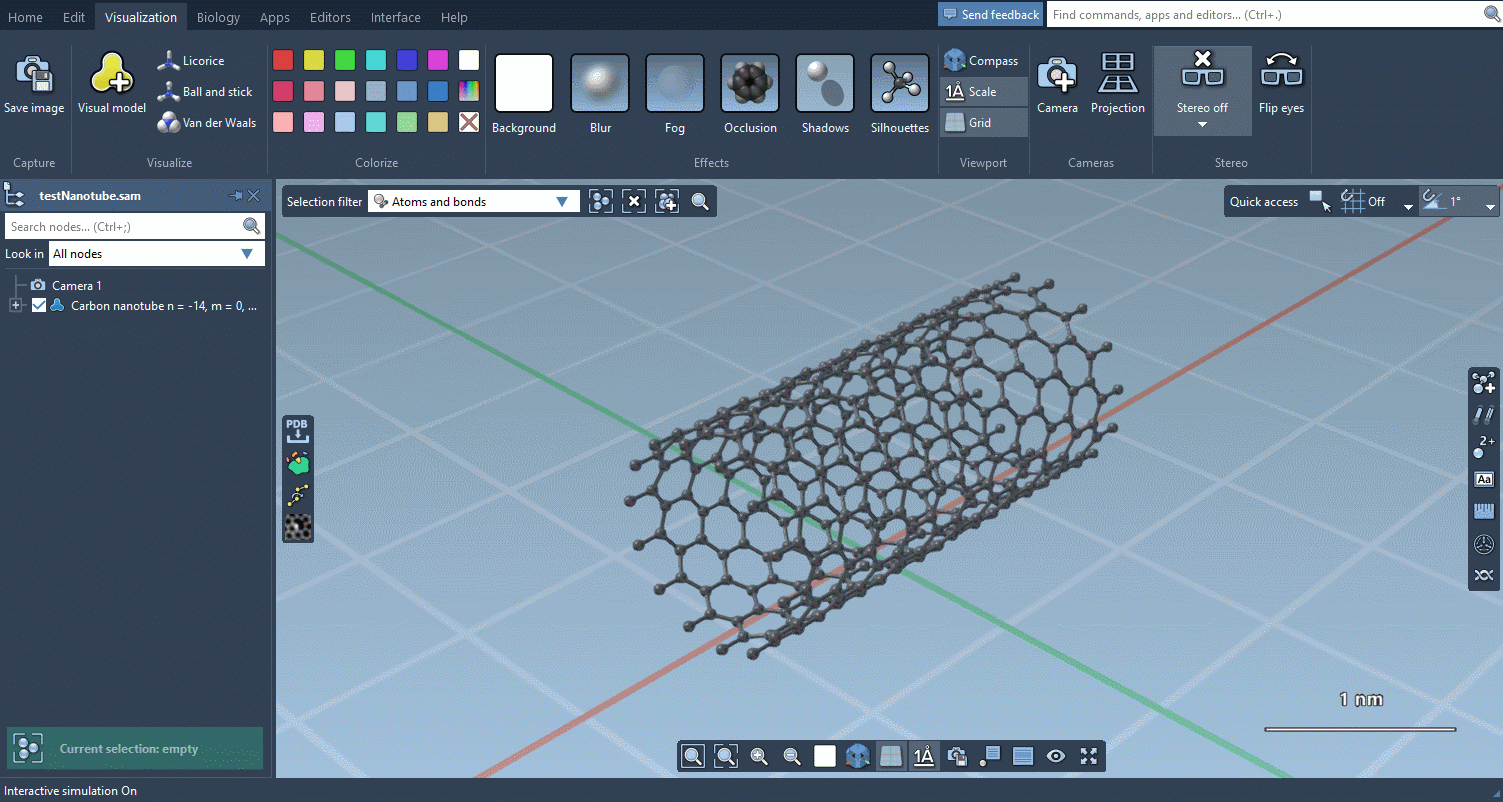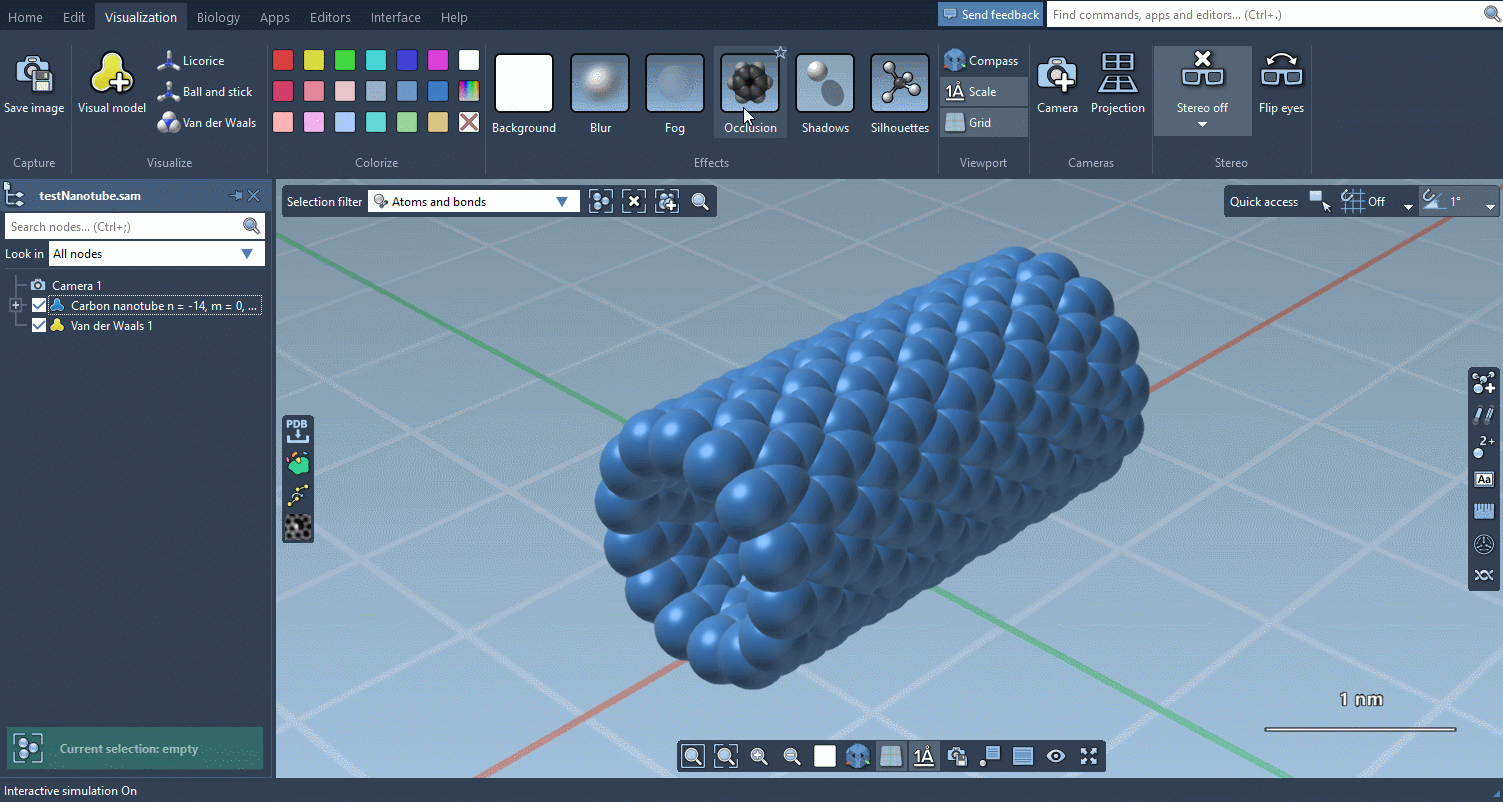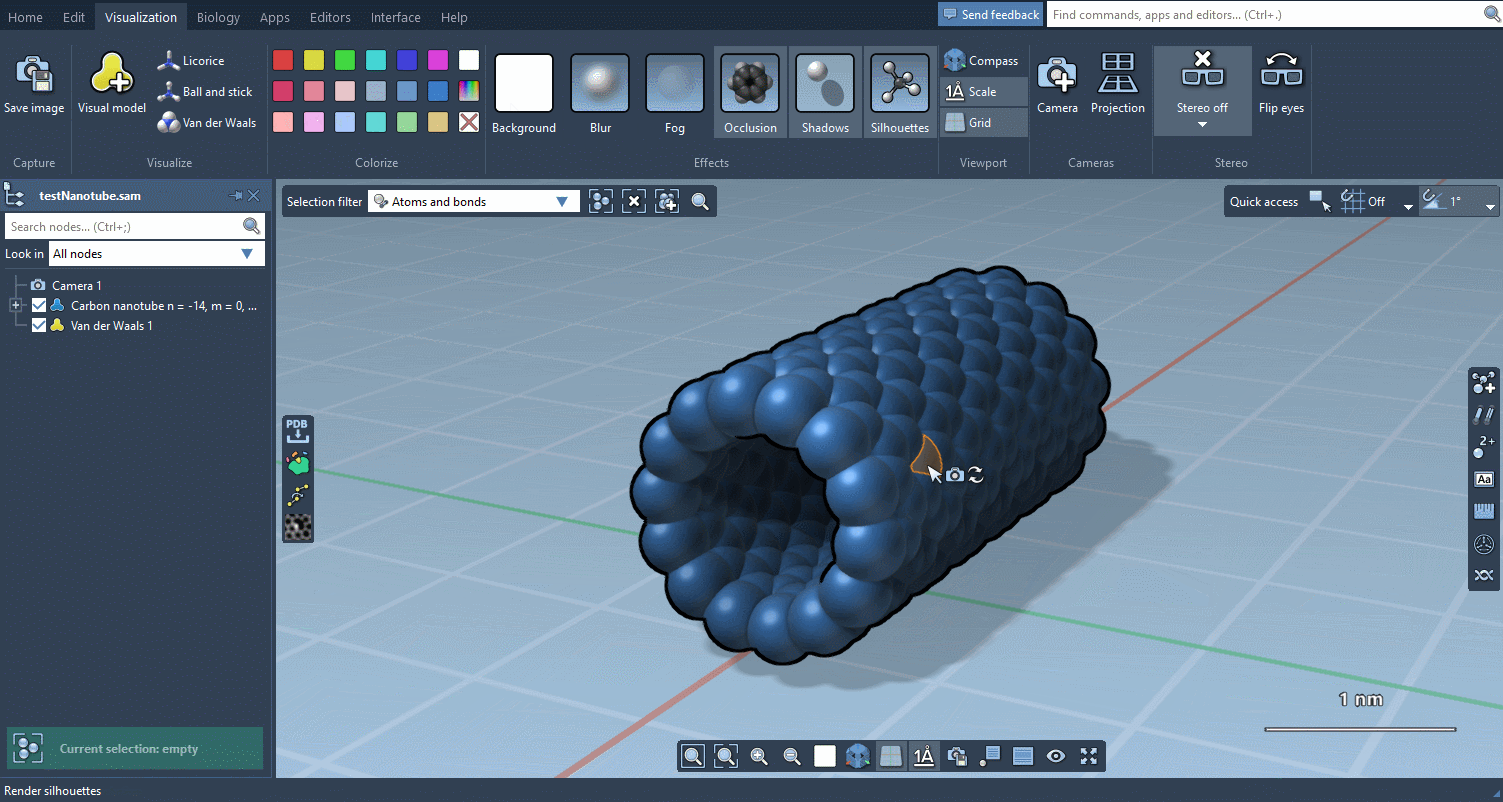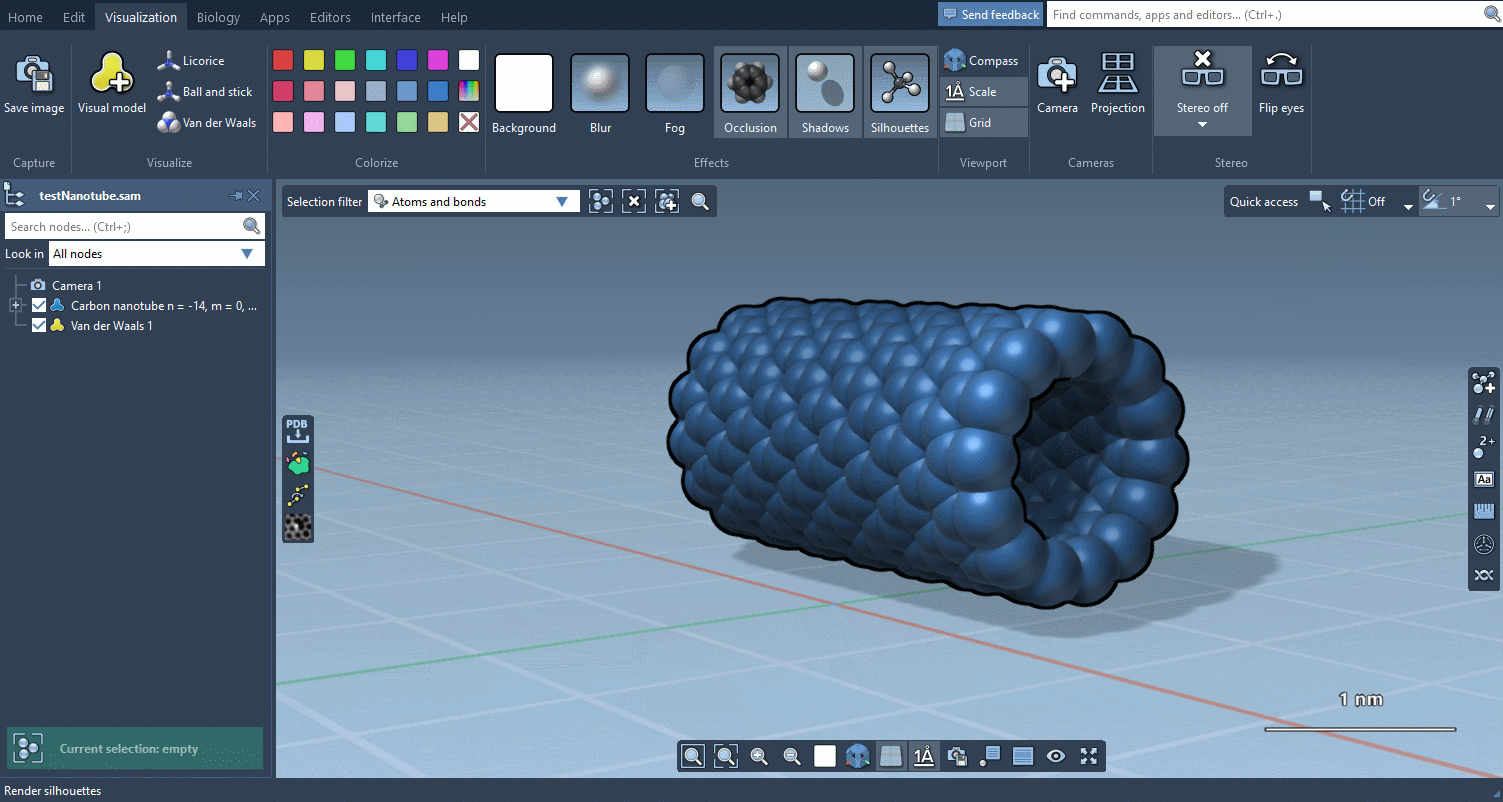
Biology menu
The Biology menu gathers specific commands for selection, visualization and per-attribute colorization of biomolecules:
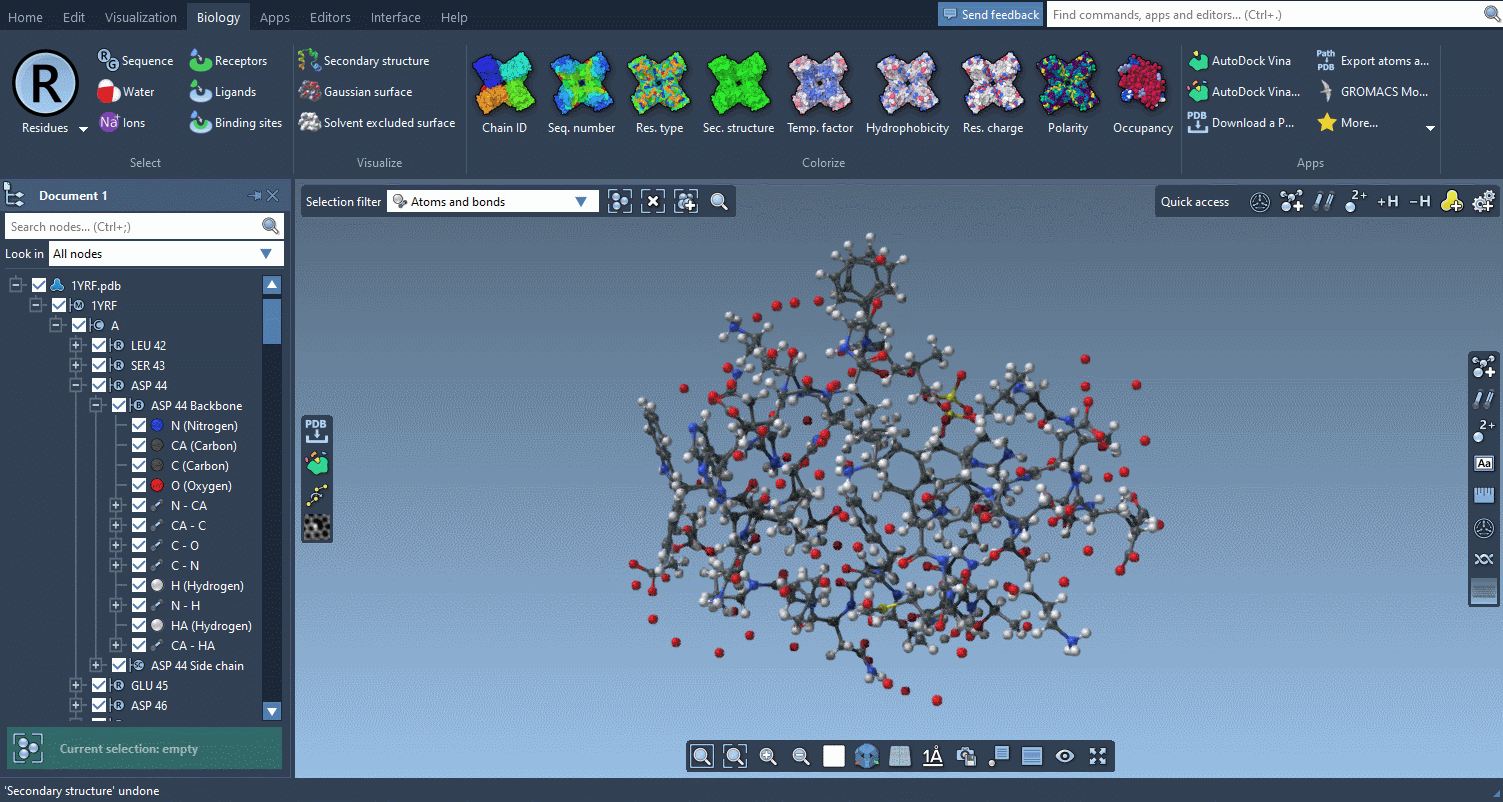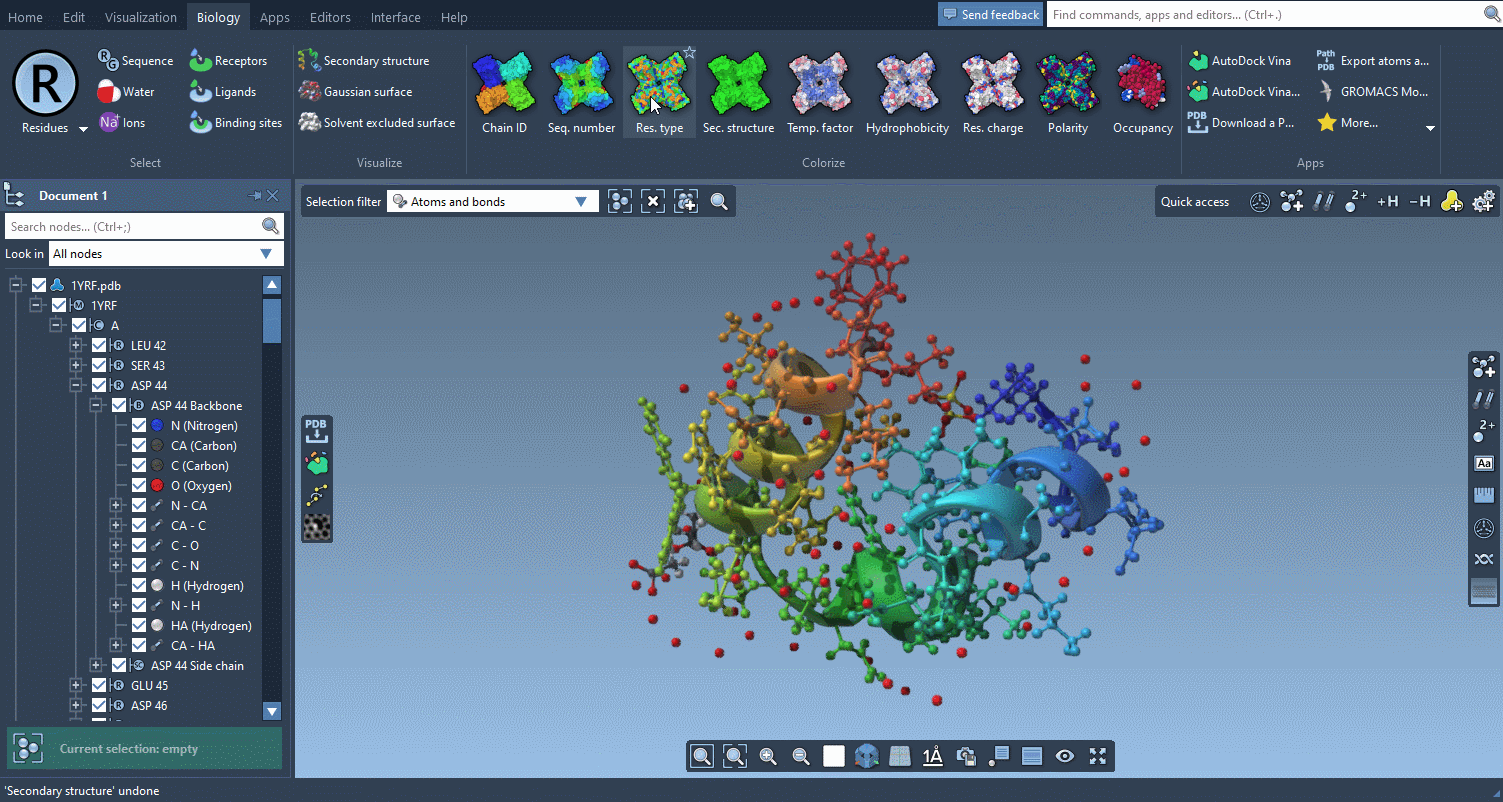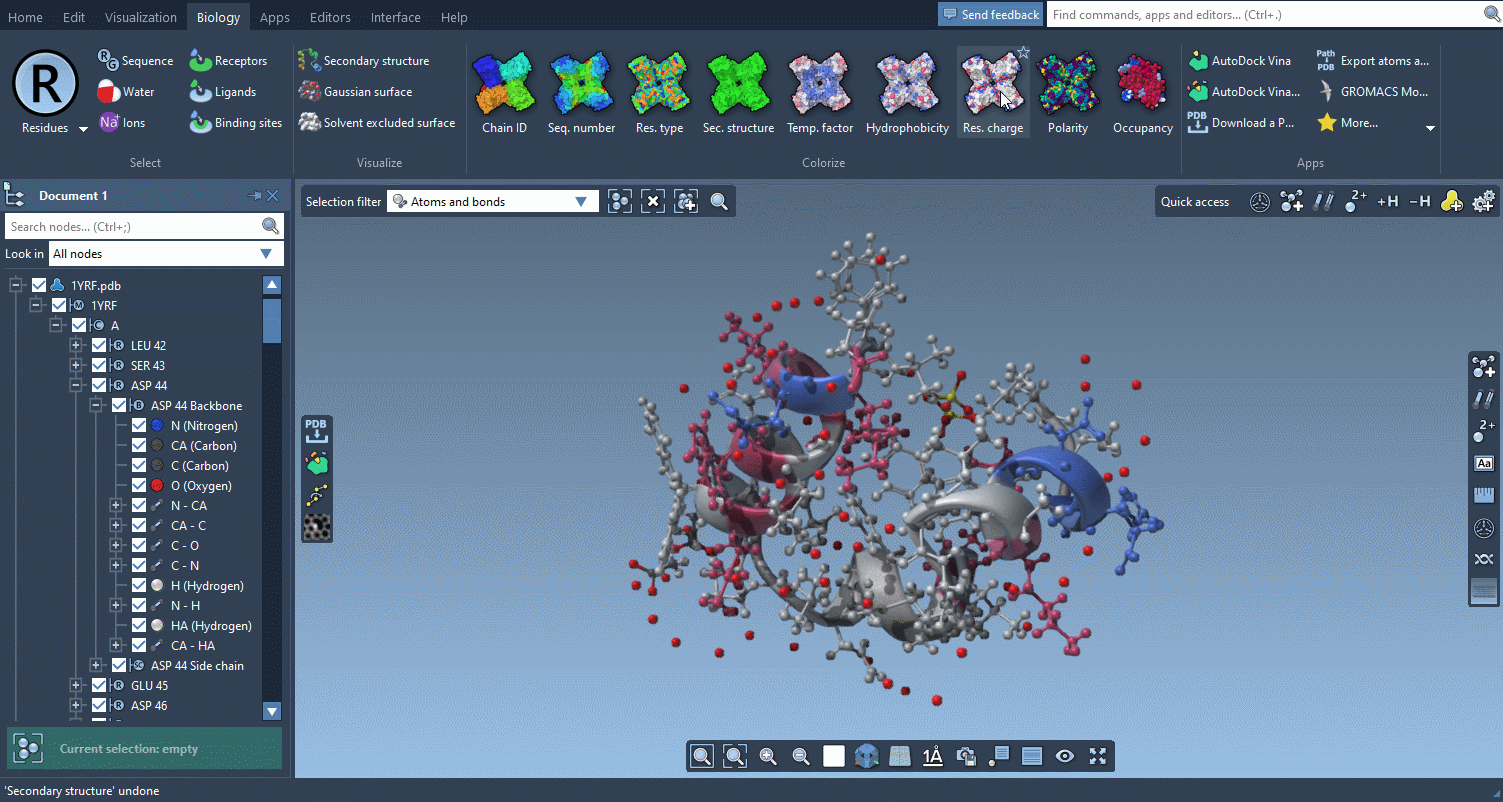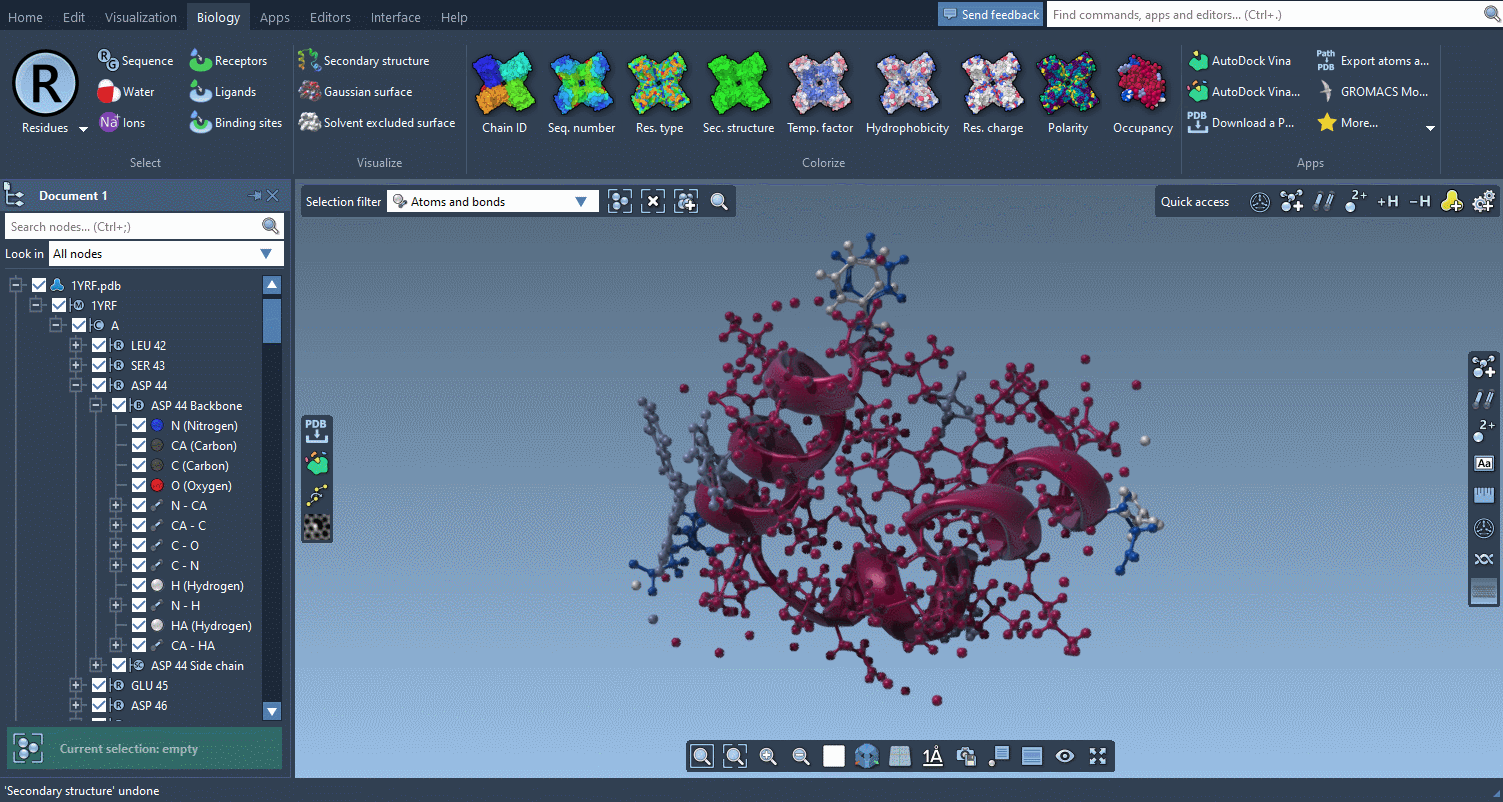
This menu is optional and can be hidden in the Preferences.
Apps menu
The Apps menu has been reorganized to show all your apps more clearly and make them more accessible. You can also use the menu to set apps as favorites and place them on the viewport overlay.
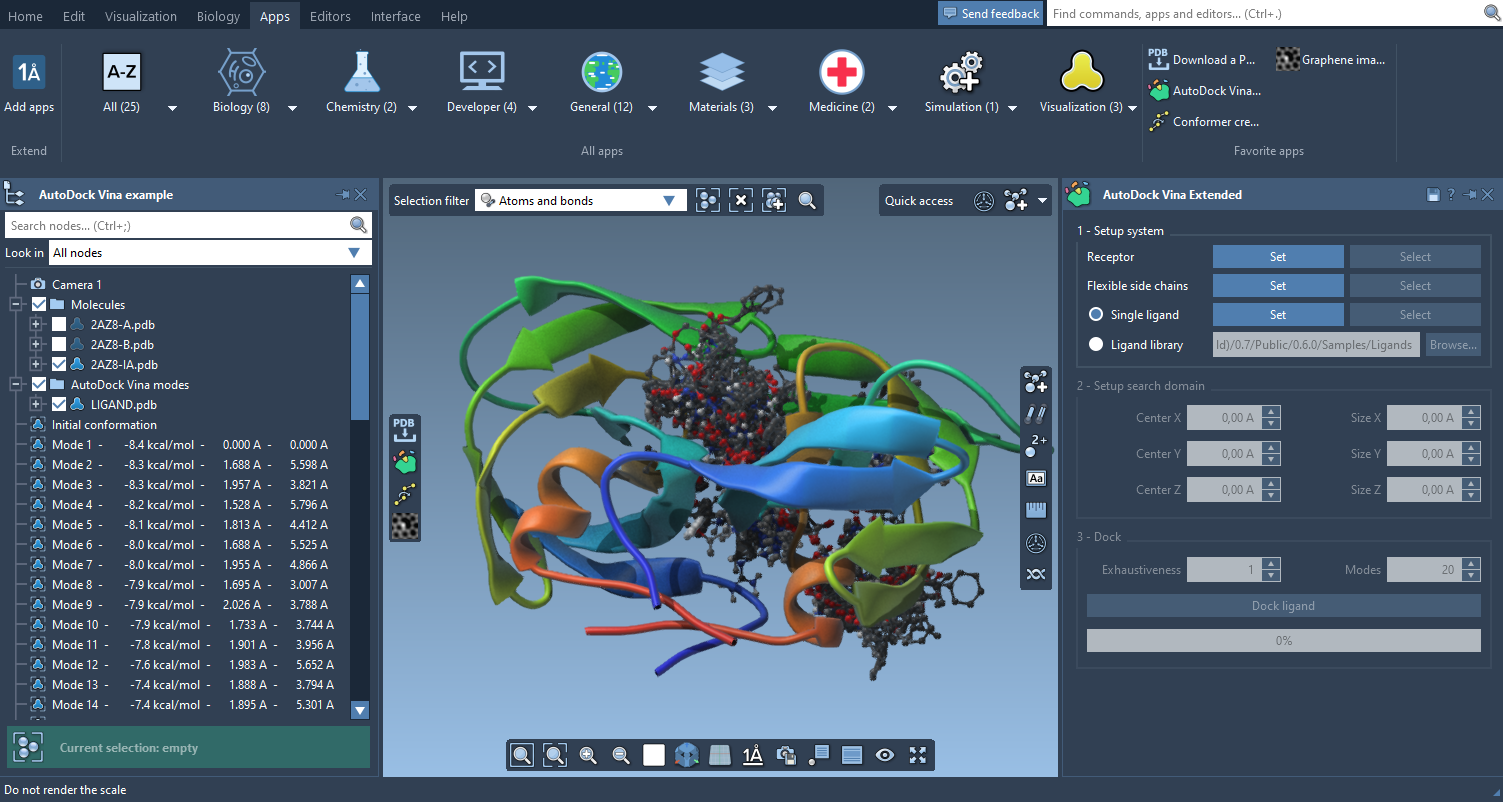
Editors menu
The new Editors menu is similar to the Apps menu and gathers all your editors. As with apps, you can set editors as favorites from this menu to make them accessible from the Viewport overlay (see below).
Command Finder
The Command Finder was improved to display icons, images, shortcuts and help:
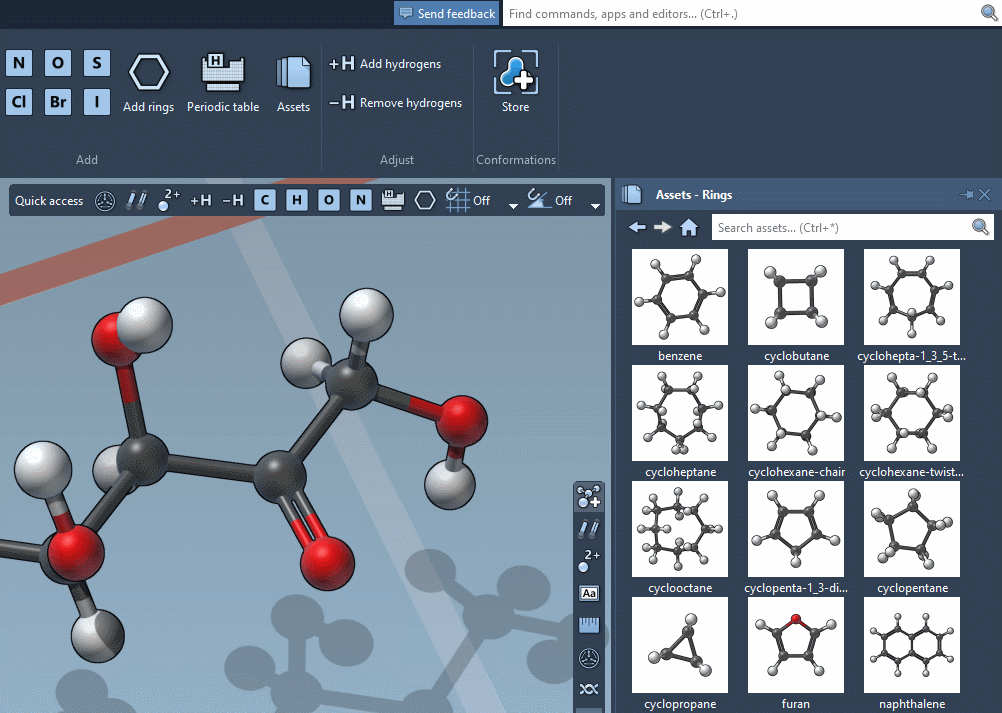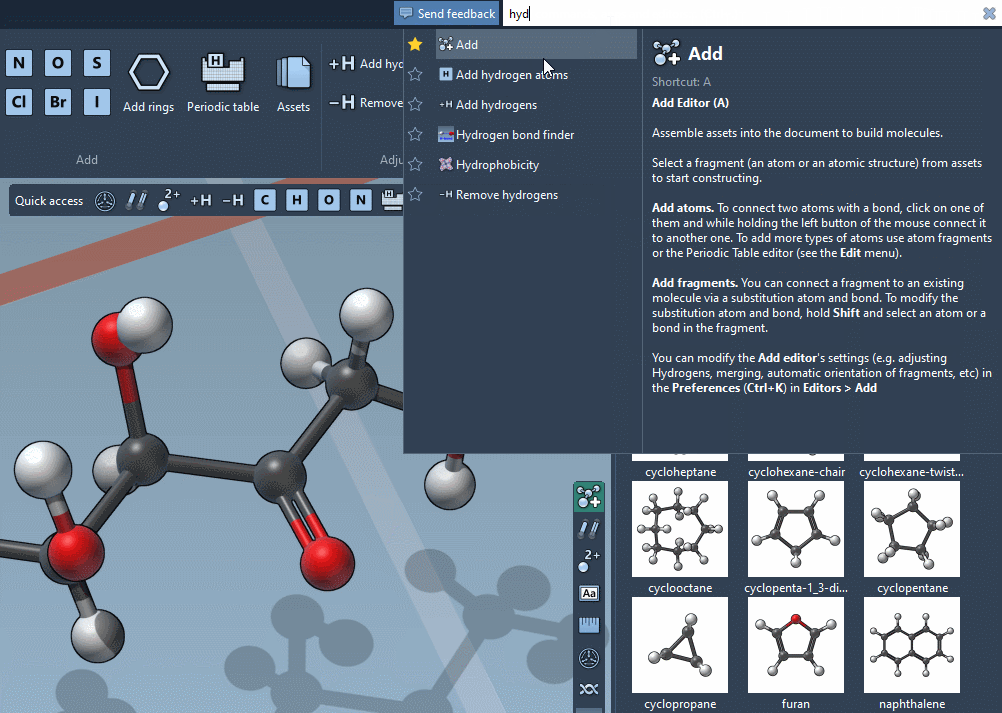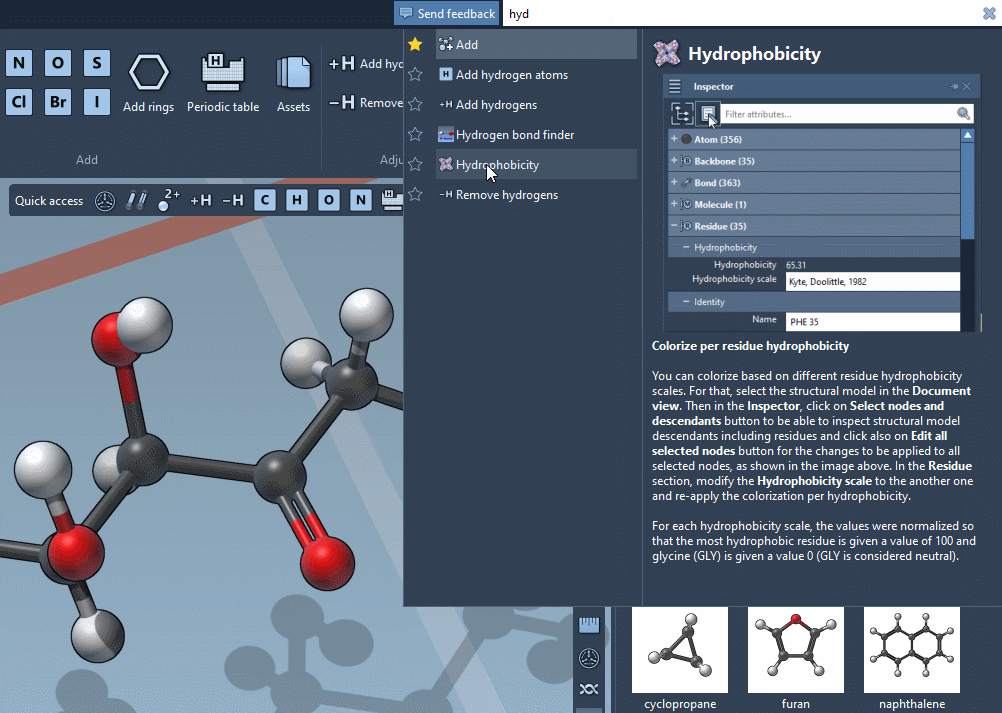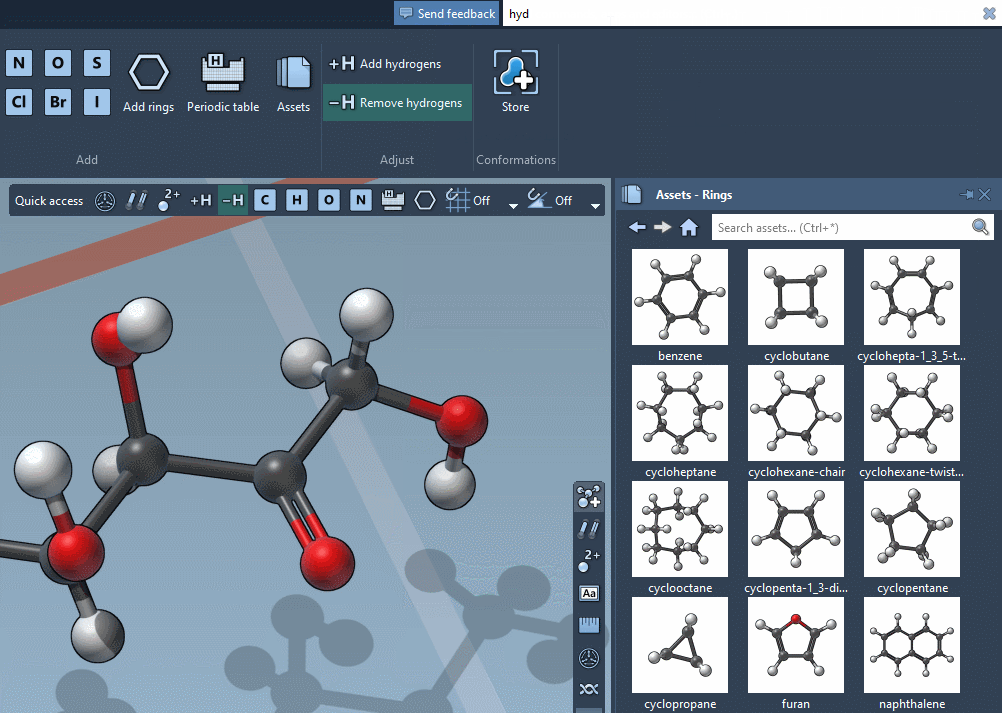
Viewport overlay
The viewport now features overlay actions to speed up your workflow:
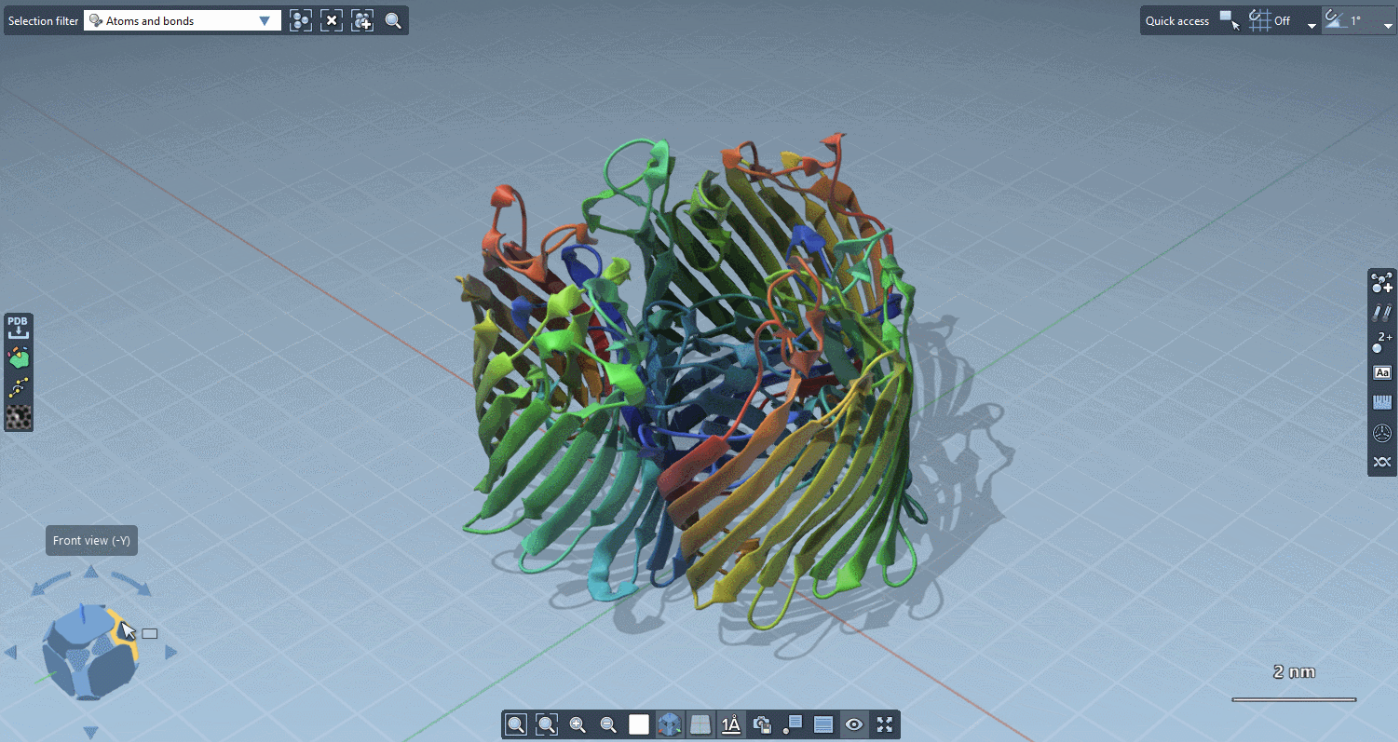
On the top-left part, the Selection filter lets you control what you select in the viewport. The left part shows your Favorite apps (you choose which apps you set as favorites). The bottom part controls the View. The right part shows your Favorite editors.Finally, the top-right part is dynamically updated to provide Quick access commands related to the active editor.
On the bottom-left part, you can use the Compass to know and set the camera orientation:
On the bottom-right part, the Scale indicates the size of the objects shown in the viewport:
Hovering nodes now displays information about them and their ascendants:
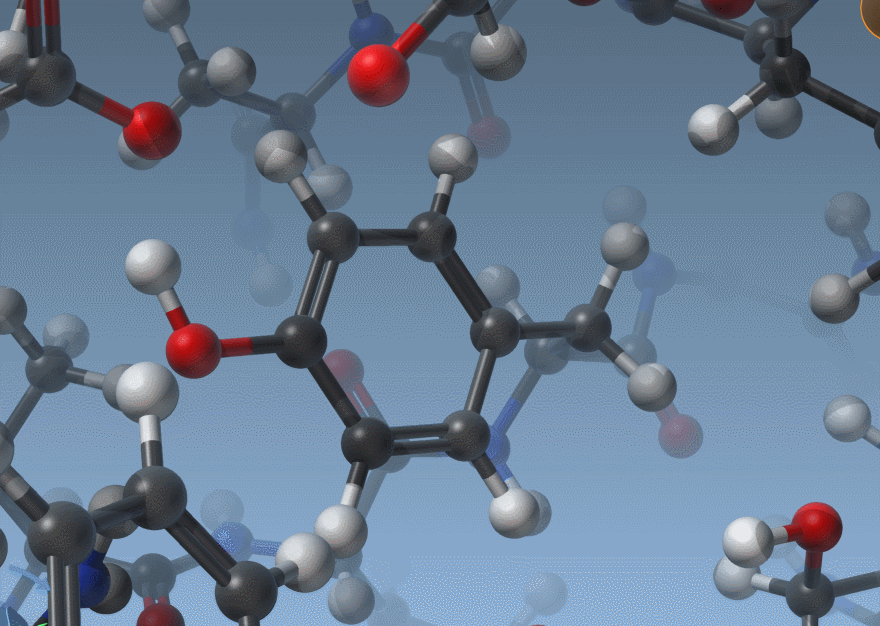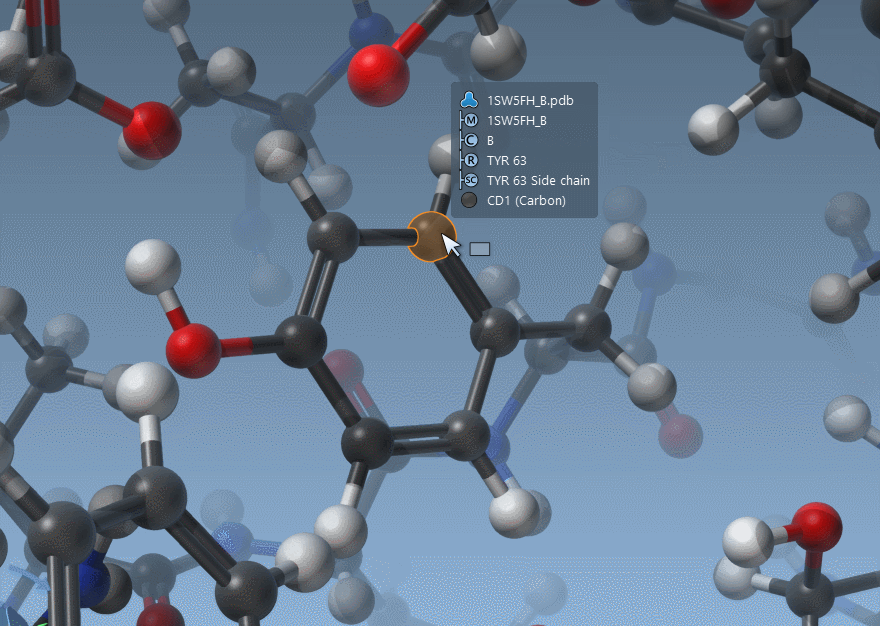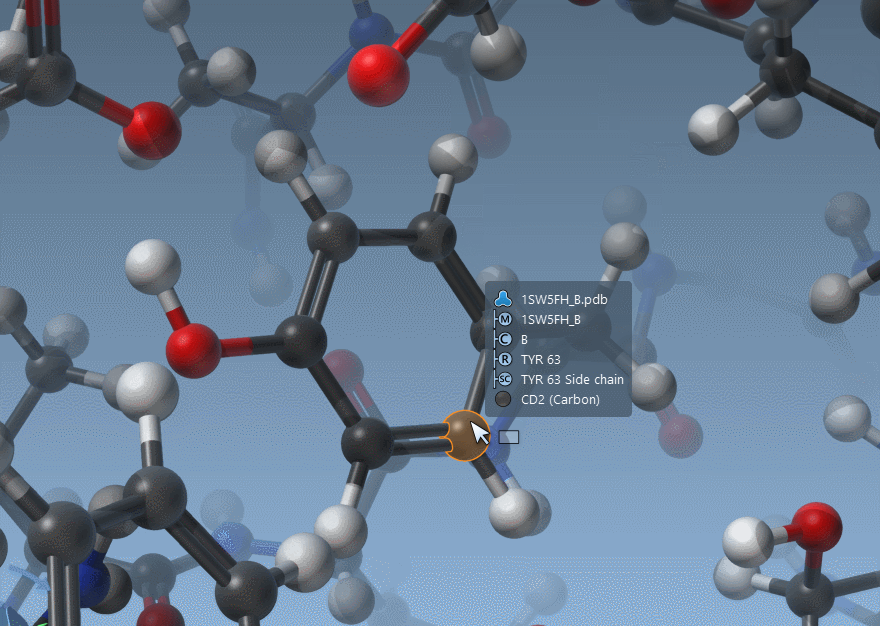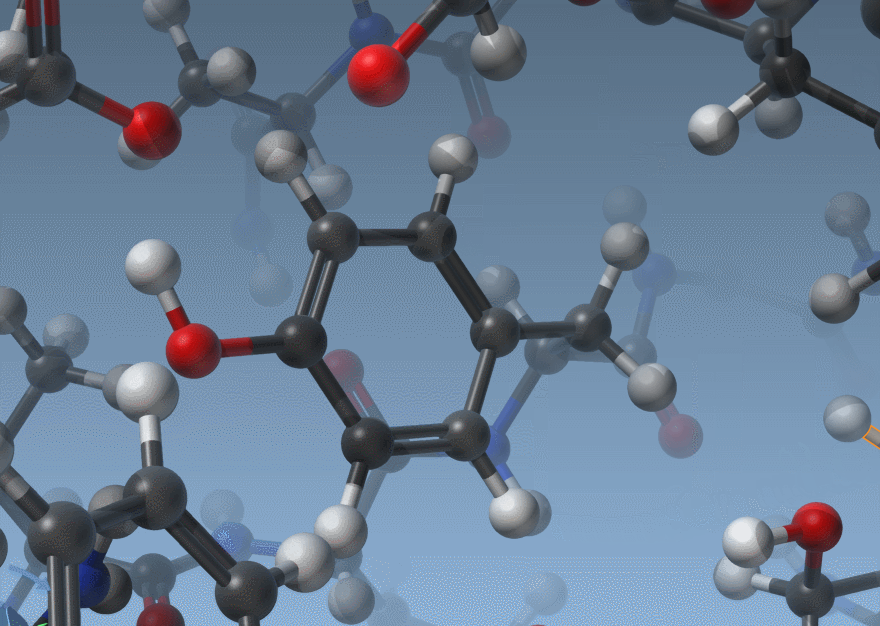
As shown above, SAMSON now has two ways to display bond orders: either through their thickness (as before) or via multiple cylinders (new version).
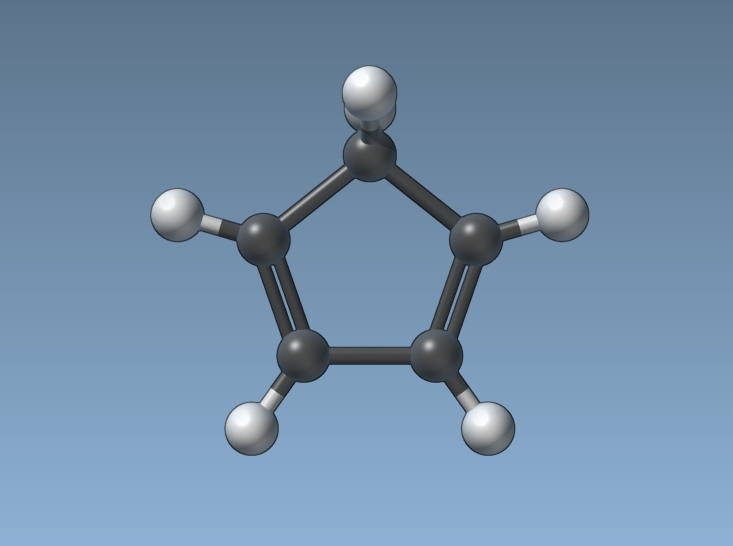
Some of you are doing simulations that involve continuous bond orders, so we decided to make rendering bond orders continuous as well:

Try it yourself: select a bond and change its order in the Inspector (Ctrl / Cmd + 2).
An easier learning curve
Interactive tutorials
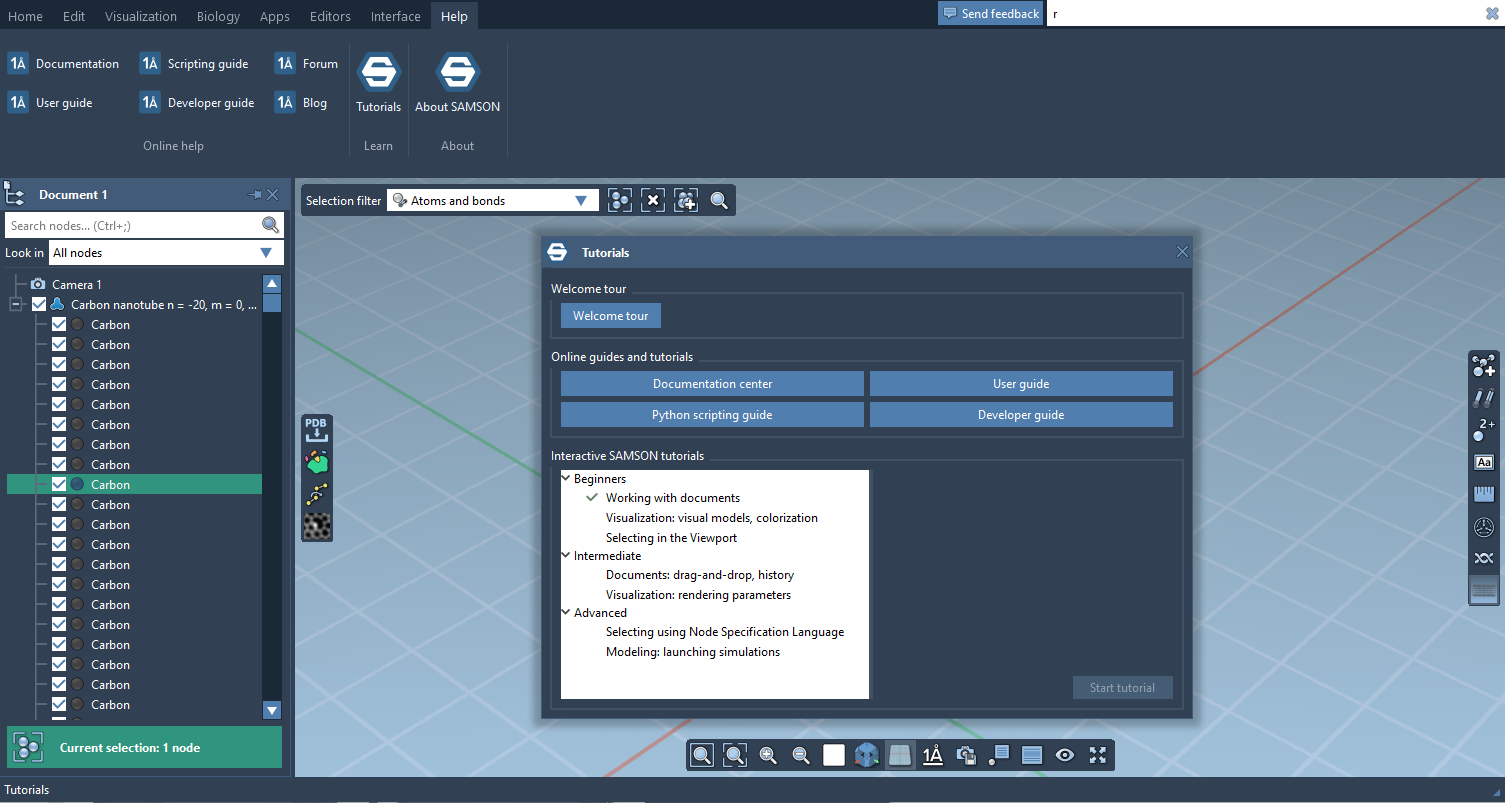
SAMSON now contains step-by-step Interactive tutorials that guide you through SAMSON’s features at your own pace.
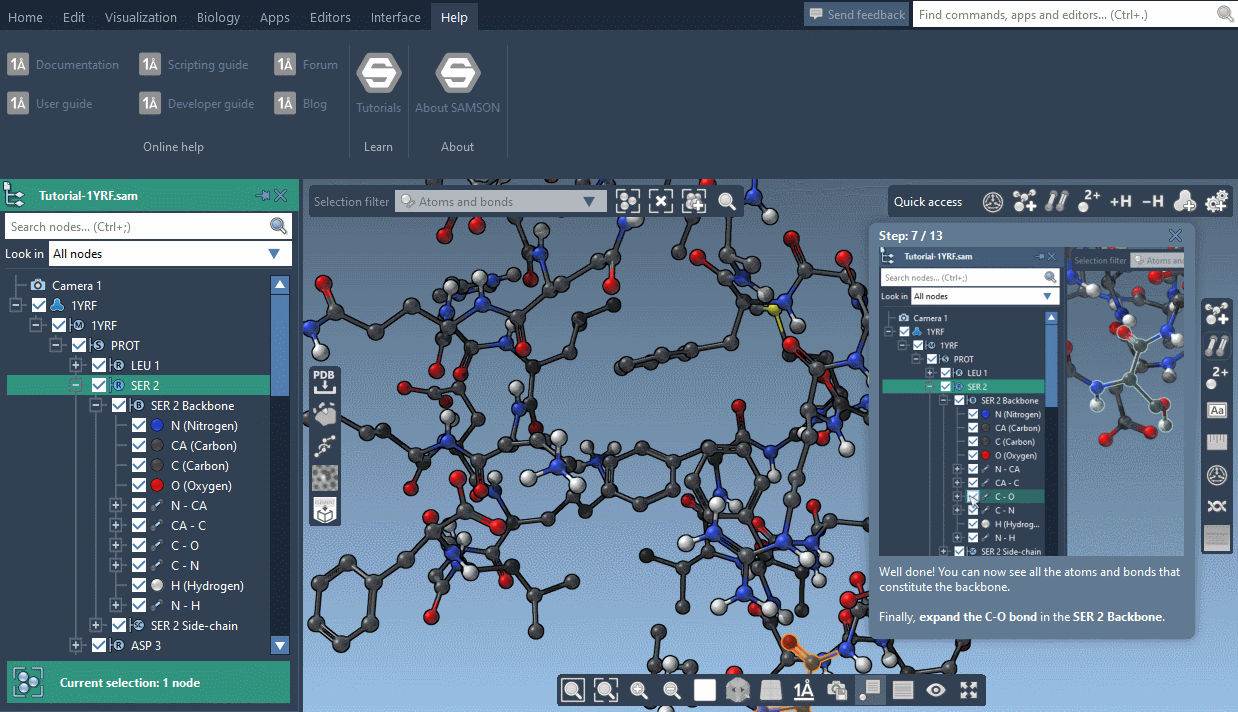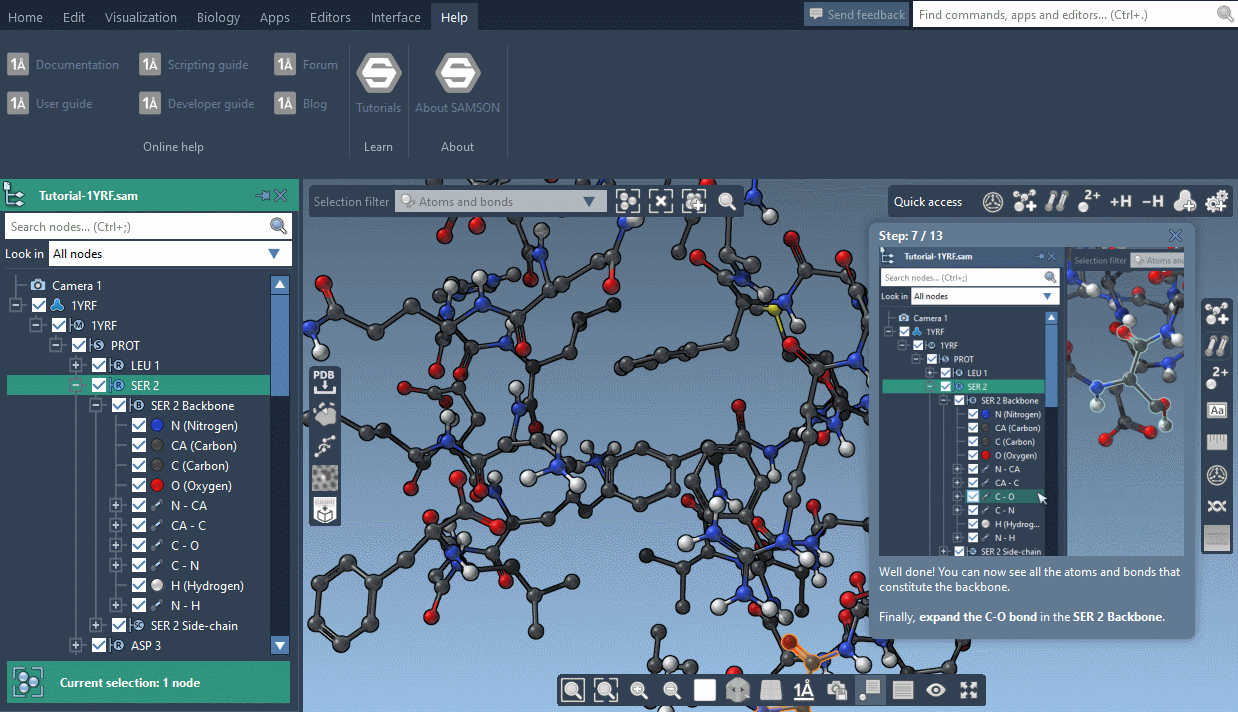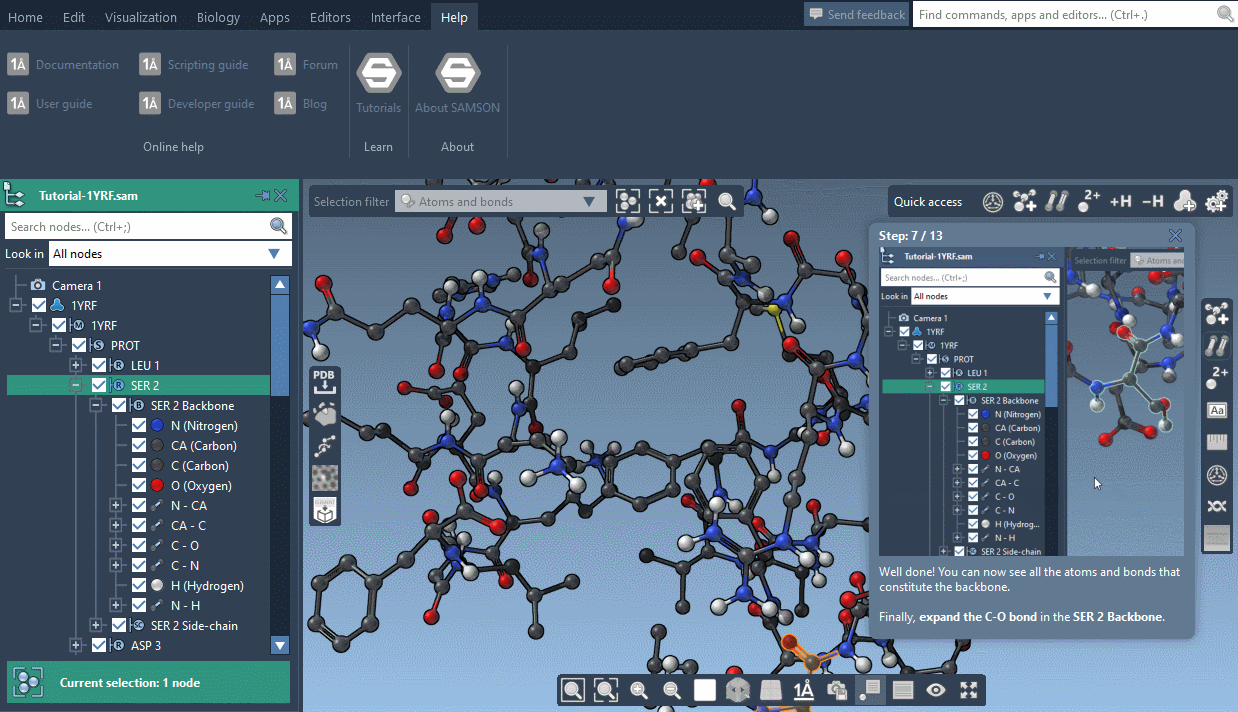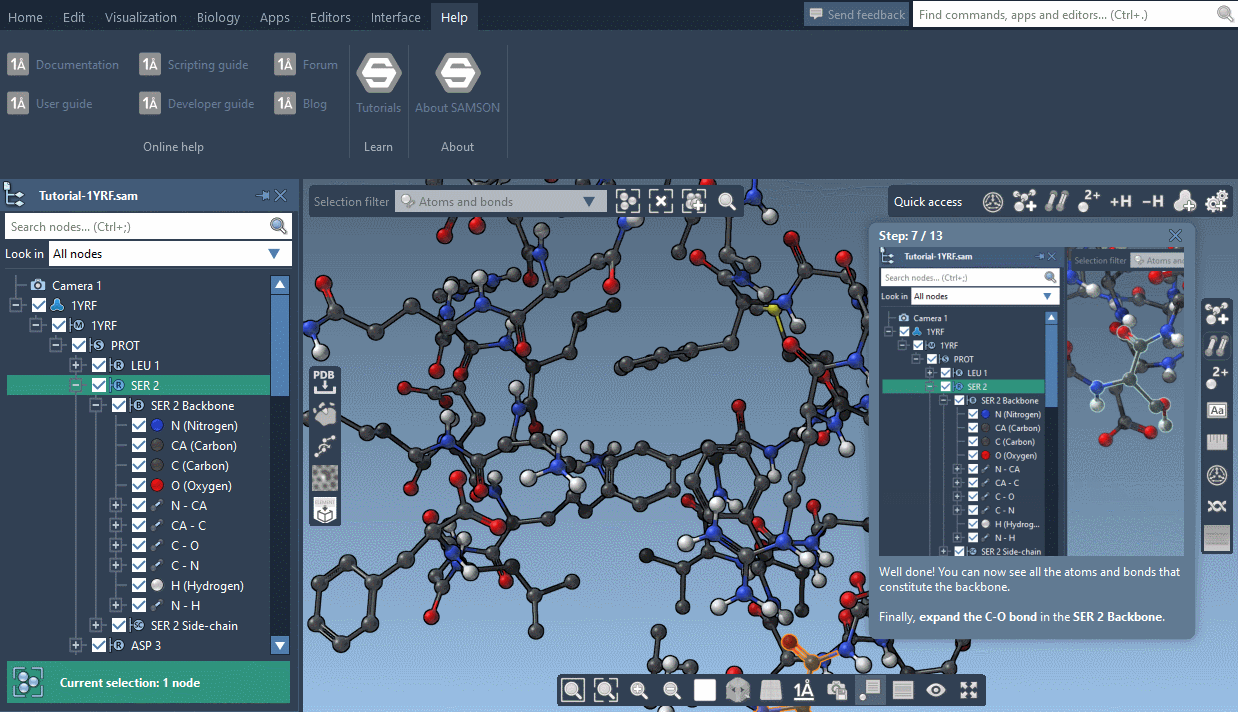
At the moment, seven tutorials are already included (totalling dozens of steps), but more are coming, and we’ll make it possible for developers to make their own and create new learning experiences, either about their own modules or about molecular science in general.
Contextual tips
The technology we developed for offering interactive tutorials is also used to provide Contextual tips which appear when they’re most needed.
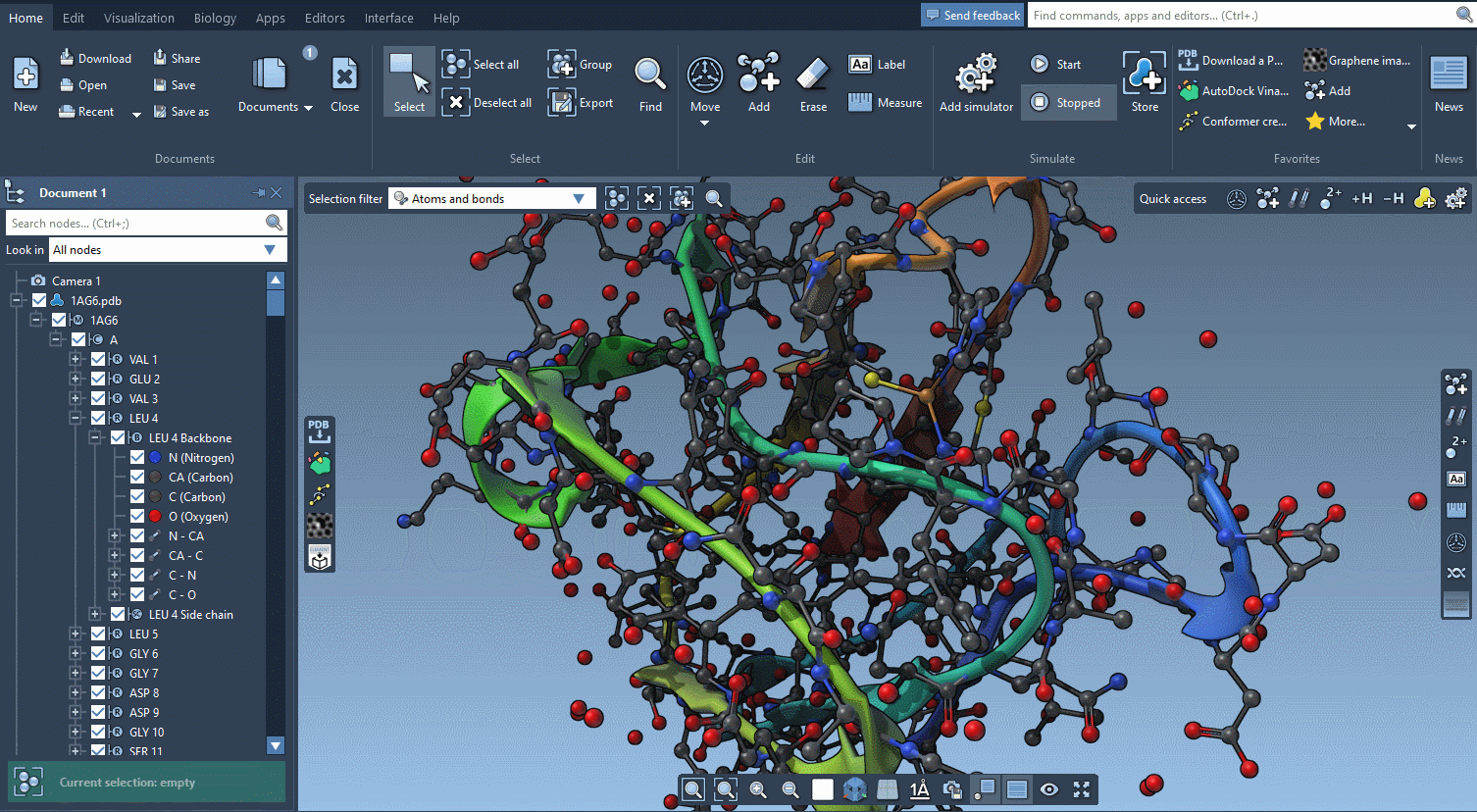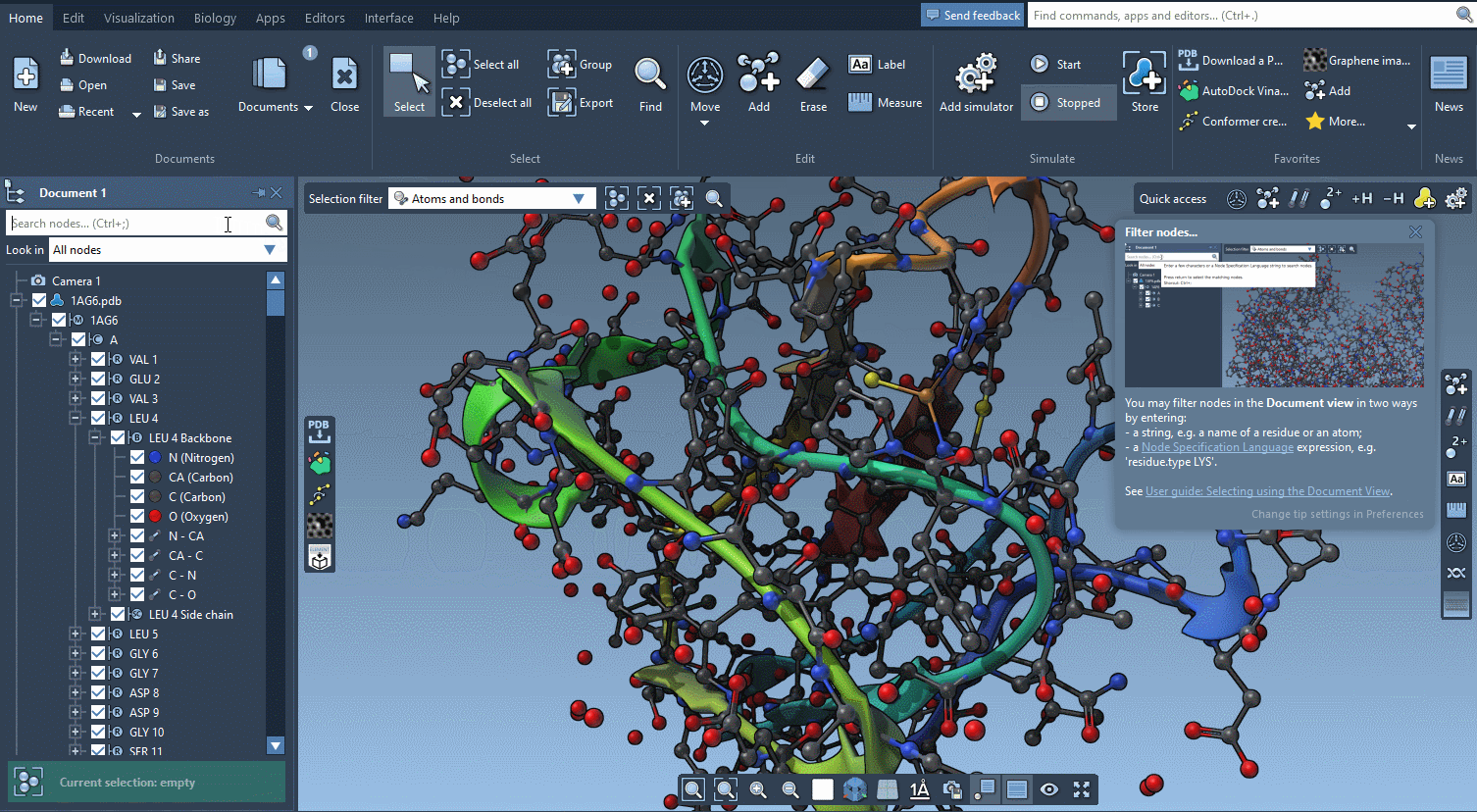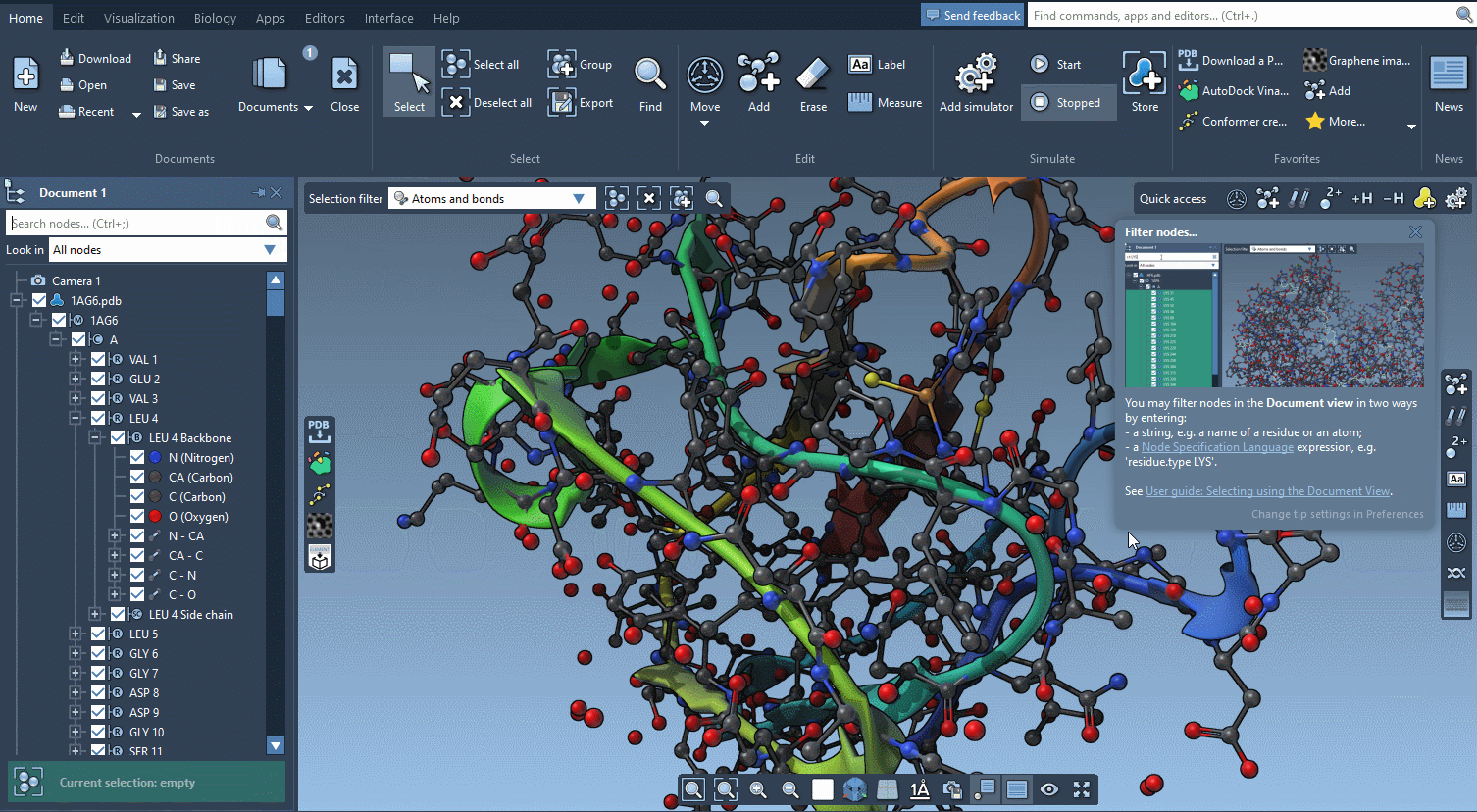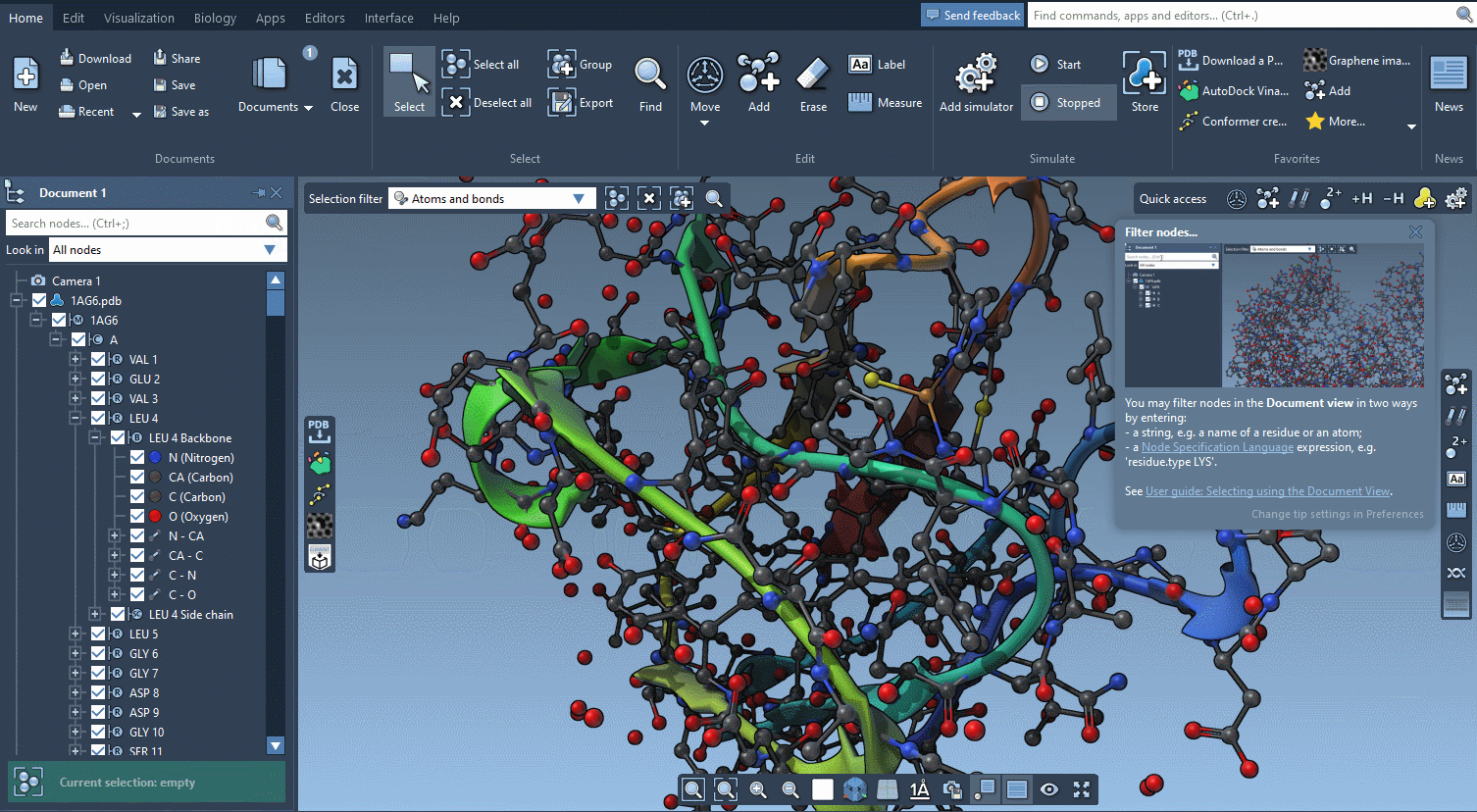
You control in the Preferences the frequency at which these tips are displayed.
And more
This is just an overview, and SAMSON 2020 includes even more new features and improvements, including for example:
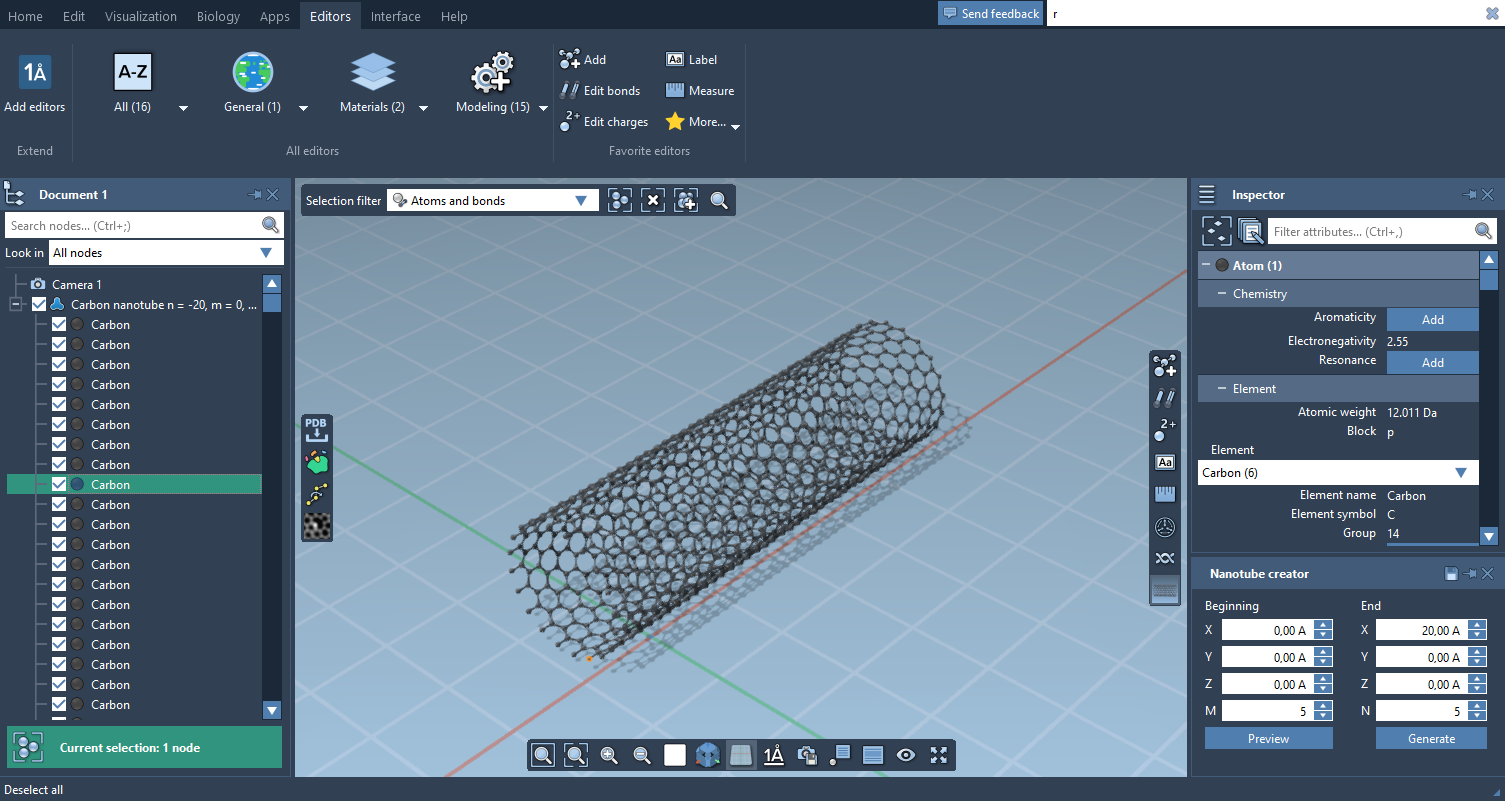
- The Node Filter can now be used to search All nodes, Visible nodes and/or Selected nodes, and it provides examples of search strings
- The Document view is orders of magnitude faster than in the previous version
- The Node Finder has been simplified to make it easier to learn SAMSON’s Node Specification Language
- The SAMSON API has been upgraded to expose the new functionalities of this release and let developers create fantastic molecular modeling experiences that they can distribute on SAMSON Connect.
To find out more, sign up on SAMSON Connect (it’s free!) and download SAMSON now!
If you have any questions or feedback, please use the SAMSON forum.





