Loading molecules#
Use this page to open local structures, fetch molecules, understand how SAMSON documents store data, and embed related files.
Start here after installation when you are ready to bring real content into SAMSON, whether that content comes from local files, fetched structures, and embedded project material.
When to read this#
Read this after Moving around, when you can navigate the viewport and recognize the Document view. After loading a structure, most workflows continue with selection, inspection, visualization, editing, simulation, or sharing.
Open local files and fetch structures#
To load a molecule use Home > File > Open or the Ctrl+O shortcut (O as in Open) on Windows and Linux or Cmd+O on Mac. To open a recent file, follow Home > File > Recent, where you will find a list of the recently opened files.
SAMSON will use an appropriate importer to load a file based on its format. By default, SAMSON has importers and exporters for multiple file formats (see the list of supported formats). If you want additional importers, go to the Marketplace on SAMSON Connect and check for other importers (see SAMSON Connect - Marketplace).
Depending on the importer and the file format you might need to specify additional parameters used by the importer, as shown below. The first time you use an importer its parameters are set by default. Once you modify them, they will be saved for the next use.
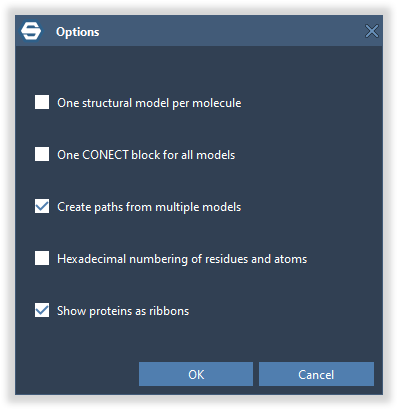
Press OK or Enter and the file will be imported into SAMSON. You will see it in the Document view and in the viewport.
Molecular structures can also be downloaded using specific Apps, such as Fetch Structures (Home > Fetch), which lets you download files (in PDB, mmCIF/PDBx, MMTF formats) from the RCSB Protein Data Bank.
Documents#
Loaded or created molecules, files, and folders are added in the active document. You can see the document in the Document view, which you can open by clicking Interface > Document view (1).
- , : Ctrl+1, : Cmd+1
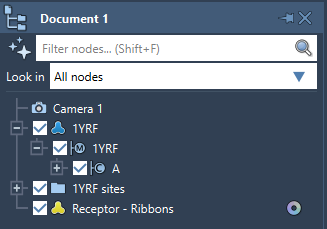
The Document view shows the data graph of the active document. Documents store all information about atoms, bonds, molecules, models, simulators, and more.
You can simultaneously have several documents open in SAMSON, however, only one document is active at any given time: the one you see in the Document view. Having several documents is useful, for example, when you want to do different tasks with different molecules or copy structures from one document to another. To switch between documents, click on the Documents list in the top-left corner of the menu, or Home > Documents (1). You can also see there the number of opened documents.
- , : Ctrl+Tab and Ctrl+Shift+Tab
: Cmd+Tab and Cmd+Shift+Tab
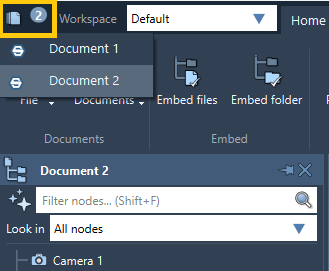
To create a new document, follow Home > File > New (1).
- , : Ctrl+N, : Cmd+N
To open recently opened documents, use Home > File > Recent.
See Document view, document, and node types for more details.
Embedding files and folders#
SAMSON documents can contain not only molecular models. They enable Universal File Embedding and can embed Python scripts and any number of files and folders. This makes it possible to embed Python scripts and whole Python apps (e.g., machine learning apps), research papers, images, data, and much more.
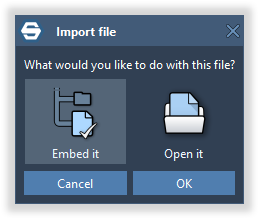
To embed files and folders within a document, drag and drop them in SAMSON and you will be asked whether you would like to embed them, or use Home > Embed files or Home > Embed folders.
Folders and files are stored within the document, making the document self-contained, so you can transfer documents between computers and share documents.
Next step#
- Continue with Work with structures if you want to select, inspect, or measure what you opened.
- Continue with Building molecules if your next goal is editing or constructing a system.
- Continue with Share documents if you need to share documents on SAMSON Connect or download shared documents.
- Continue with Collaboration if you need to manage access rights, groups, or shared documents on SAMSON Connect.