Protein Alignment#
Learn how to use SAMSON's Protein Aligner extension to align/superimpose proteins at both the sequence and structural levels - an essential tool in comparative protein-modeling and structure-based molecular design.
Watch a quick demo:
Note
This extension is included with all the plans and can be accessed via Home > Align ![]() .
.
Video tutorial#
You can learn how to use the Protein Aligner from the following extract of the SAMSON 2025 webinar:
Why Align Proteins?#
- Identify conserved residues that drive function or ligand binding.
- Compare conformations across species or mutants.
- Assist in building accurate homology models for downstream molecular design workflows.
Fetch Example Proteins#
We'll align two hemoglobins from different organisms: 1DLW and 1RTX.
- Open Home > Fetch.
- Input
1DLW 1RTXin PDB or PDB (mmCIF). - Click the corresponding Load button.

Clean Up the System First?
Use Home > Prepare to strip water, ligands, or alternate locations. See the Protein Preparation & Validation guide for details.
Launch Protein Aligner#
Click Home > Align ![]() to open the Protein Aligner.
to open the Protein Aligner.

Sequence Alignment#
You can align sequences in two modes:
- Align sequences (by structure) - aligns one model to another (per structural model).
- Align sequences (by chain) - aligns individual chains (there can be multiple chains per model); useful for oligomers.
Click either Align sequences button to generate the alignment:

To see if the corresponding amino acids have the same properties, you can toggle amino acid property options via ![]() Highlight residues. For example, on the image below, we can see the comparison of two sequences based on the similarity (shows the conserved residues) and polarity of amino acid residues:
Highlight residues. For example, on the image below, we can see the comparison of two sequences based on the similarity (shows the conserved residues) and polarity of amino acid residues:

Note
If a residue matches multiple selected properties, its color blends the corresponding shades, as can be seen from the image above.
Interacting with the Alignment#
-
Hover over a residue in the sequences to see its name and ID and highlight it in the viewport:
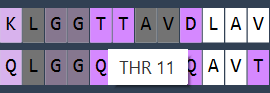
-
Click on a residue - selects that residue and its aligned partners.
- Hold Shift to select multiple consecutive residues.
- Hold Ctrl/Cmd to select non-consecutive residues.
- Hold Alt to remove from the selection.
To clear the selection, click anywhere in the viewport or press Ctrl/Cmd + D.

Tip
Selecting a residue in the Document view or in the viewport automatically highlights it - and all aligned residues - in the Protein Aligner.
Aligning structures#
Whole-Protein Superposition:
- Make sure no residues are selected.
- Click Align to this on the first model's row.
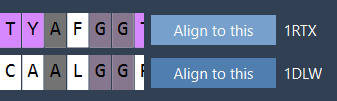
The button on the other model now reports the RMSD (e.g., 3.27 Å):
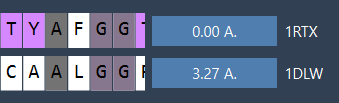

Your proteins are now superimposed:
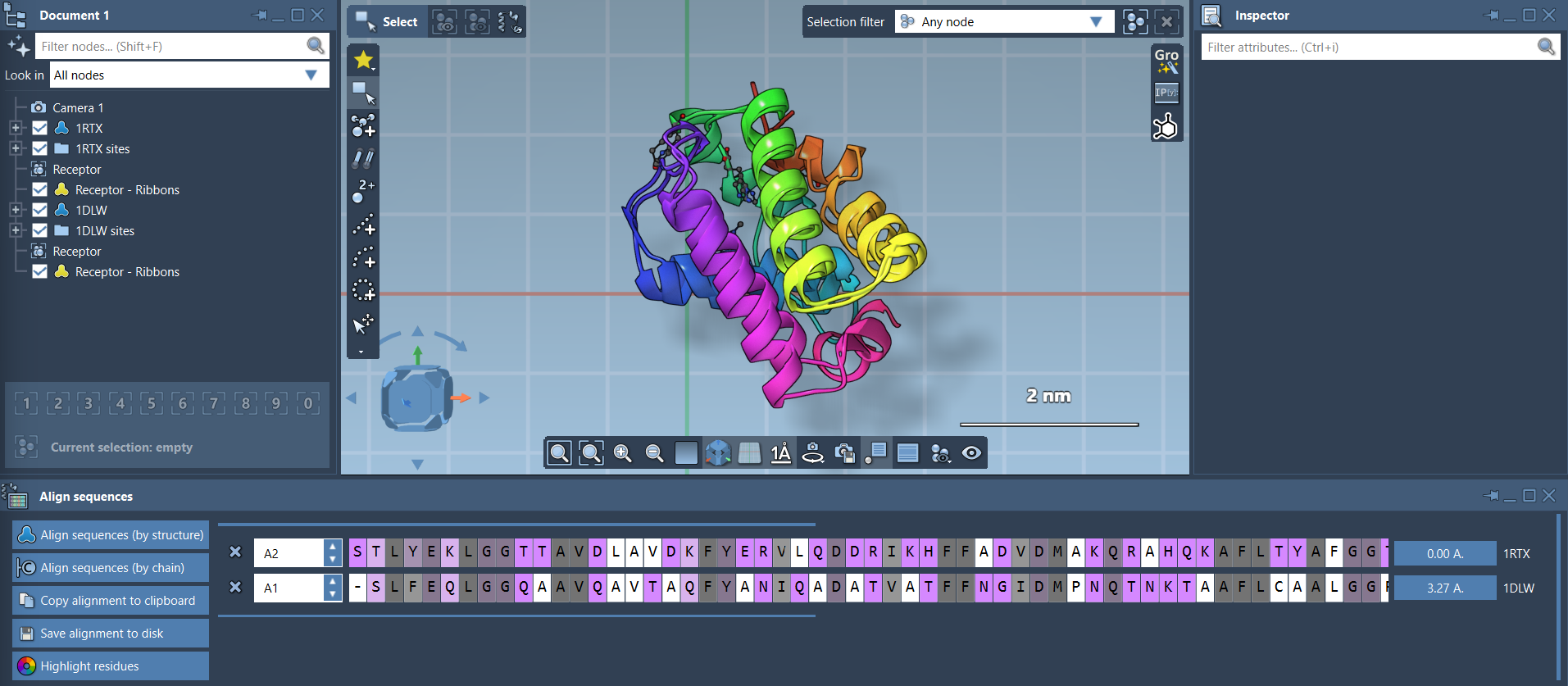
Visual Aids
If you do not have secondary structures shown, you can add them via Visualization > Visual model > Ribbons.
Create separate ribbon models per protein to color them differently. For that, select a structure first and then apply a visual model. This should help to differentiate the aligned structures.
Toggle atomistic views via checkboxes in Document view.
Region-Specific Alignment#
Sometimes only part of the protein is of interest. You can also align based only on the selected part of the structure. From the picture above, we can see that the two pink alpha-helices in the beginning of the protein sequence look very similar but are not well aligned. To align those two parts in particular, go back to Protein Aligner and select the first 20 residues in both sequences:

Click the alignment button next to the selection (e.g., the 0.0 Å button) to superimpose only those residues.

Next Steps#
- Export the alignment for homology modeling.
- Map conserved residues onto ligand-binding sites for molecular design projects.
- Repeat with additional chains or related proteins.
Need Help?#
Have questions or feedback? Feel free to reach out via the Forum, via e-mail, via the Feedback button in SAMSON, or by directly discussing with us.