Build Custom Polymers with Polymer Builder#
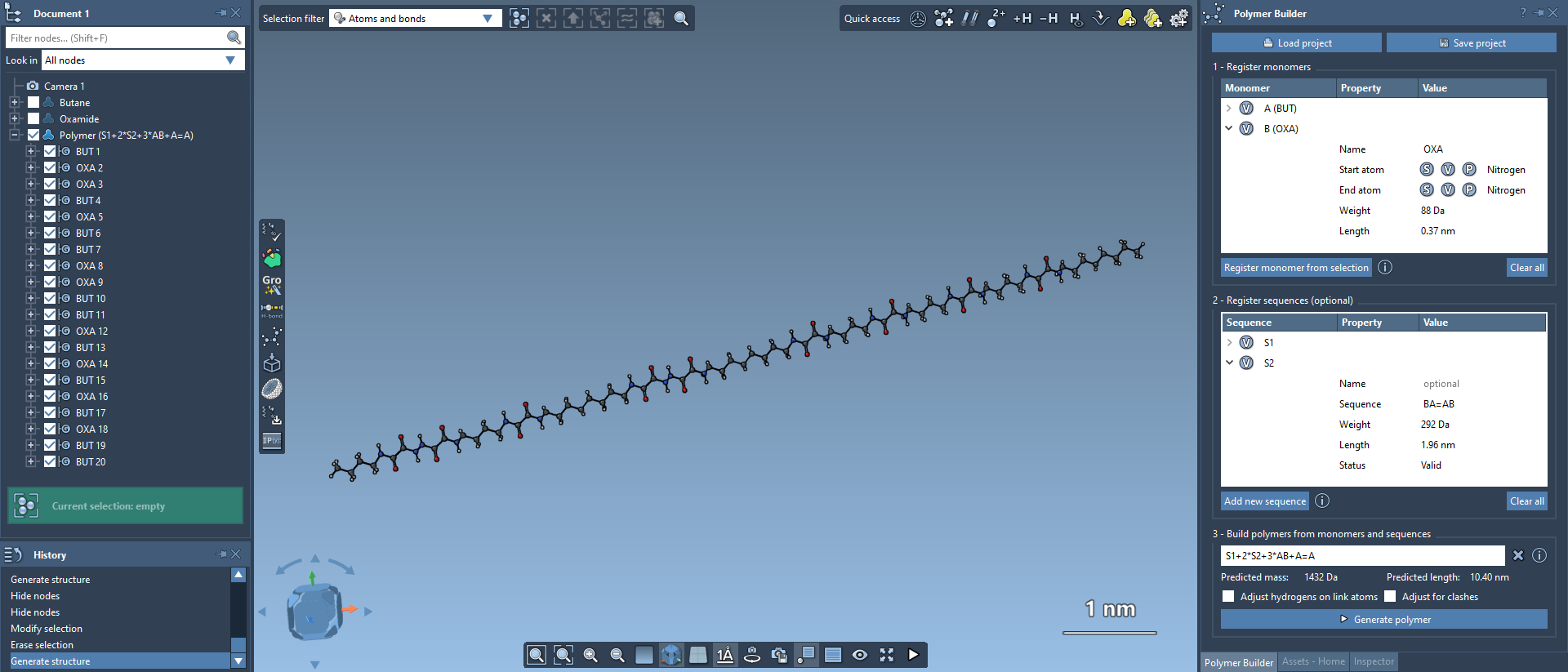
The Polymer Builder extension in SAMSON lets you build flexible, complex polymers using custom monomers or sequences. It's ideal for constructing synthetic chains, biopolymers, and custom molecular scaffolds for molecular design, simulation, or property prediction.
By the end of the tutorial, you will know how to register monomers, define reusable sequences, and generate a polymer directly inside SAMSON.
Why Use Polymer Builder?#
- Create linear or branched polymers using real chemical fragments.
- Mix & match monomers and sequences.
- Control bond types (single, double, triple).
- Automatically detect connection atoms and avoid clashes.
- Can be used for polymer design, coarse-grained modeling, etc.
What you will learn#
In this tutorial, you will learn how to assemble custom polymers from monomers and sequences with Polymer Builder in SAMSON.
Before you start#
- Log into SAMSON Connect.
- Visit the Polymer Builder Extension page and click Add.
- Restart SAMSON - your extension will be ready to use.
- If you already have candidate monomer fragments, keep them in the document so you can register them directly from the selection.
Step 1 - Launch the Polymer Builder App#
Find it in Home > Apps > Assembly > Polymer Builder or via the Find everything... (Shift+E) in the top menu of SAMSON.
Step 2 - Register New Monomers#
- Select a molecule (monomer unit) in the Document view or Viewport.
-
Click Register monomer from selection to add a new monomer based on the current selection.
- The tool automatically picks start and end atoms at opposite ends of the fragment.
-
You can override these using:
- S: Set atom from current selection (first, select a single atom in the Document view or in the Viewport).
- P: Pick atom from a list of atoms in the structure.
- V: Highlight structure or atoms.
Each monomer is automatically given a unique letter (A, B, C, ...).
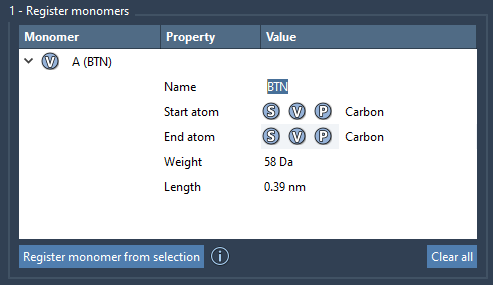
You can modify the already registered monomers right in the table and expand them if necessary.
Note
If the structure has a single residue or a structural group, then the monomer's name will be based on it, else you can provide the name if you would like this monomer to be placed in a separate structural group in the resulting polymer, i.e., having each monomer in a separate structural group.
You can see in the table the molecular weight (in Daltons) and the distance between the start and the end atoms for each monomer.
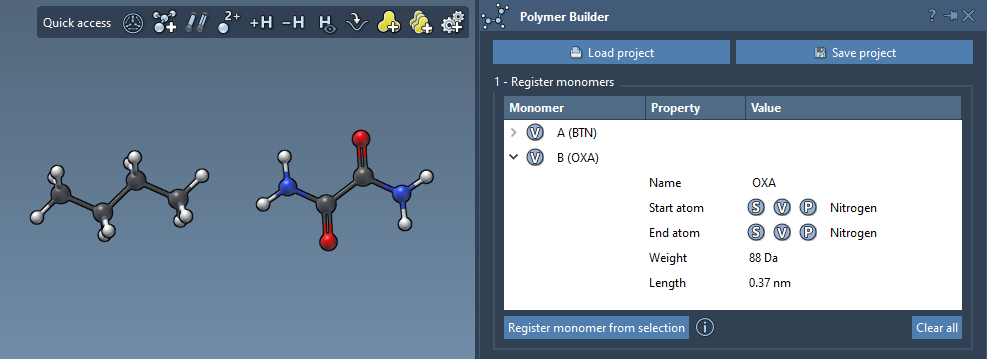
To remove a monomer from the table, right-click on it and click Delete monomer in the context menu. To clear the whole table, click Clear all.
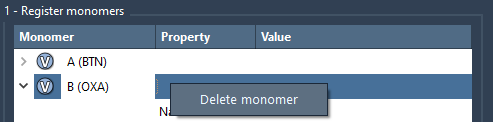
Note
If the structure of the added monomer has been modified in SAMSON (e.g., an atom has been added or removed from it) then it will be deregistered from the list.
Step 3 - Register Monomer Sequences#
If your polymer has repeating sequence patterns, you can also register such patterns as sequences of monomers:
- Click Add new sequence.
- Enter sequence using registered monomer IDs (e.g.,
ABBA).
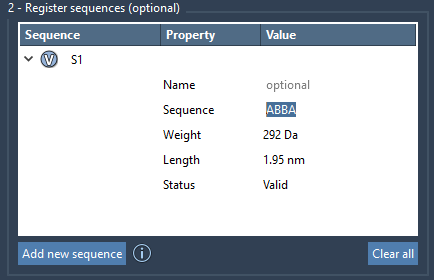
Click the associated V button to highlight the monomers used in the sequence.
If the sequence is incorrect, then the error message will be shown in the Status property; otherwise, it will be shown as Valid:
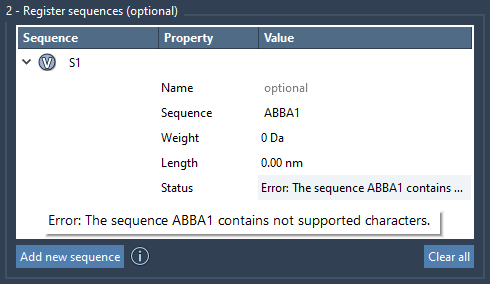
Each sequence is labeled automatically with unique identifiers (S1, S2, etc.).
You can specify the bond types between monomers. By default, the monomers are connected with a single bond. Use = for double bonds, # for triple bonds. Examples:
A=B-Aconnected by a double bond withB.A#B-Aconnected by a triple bond withB.
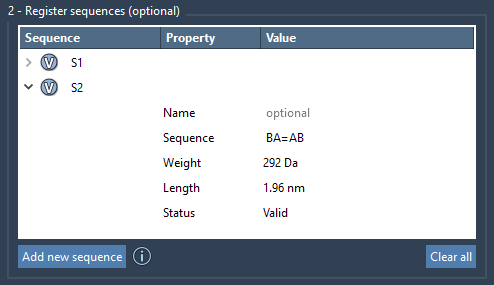
If you would like the monomers in a sequence to be arranged into a common parent structural group in a generated polymer, then assign a name in the Name property.
You can modify registered sequences of monomers (names and sequences) right in the table.
The table also shows approximate molecular weights and lengths for each sequence of monomers.
To remove a sequence from the table, right-click on it and click Delete sequence. To clear the whole table, click Clear all.
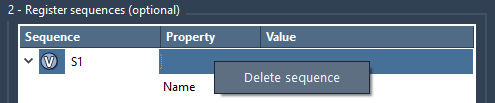
Step 4 - Generate a Polymer#
Enter a custom sequence expression combining monomers and sequences:
The sequence expression has the following format:
S1 + 2*S2 + 3*AB + AB=BA + A#B
where you can provide identifiers for both monomers and sequences of monomers, the number of repeats (e.g., 3*AB – 3 repeats of AB, i.e., ABABAB), and the bond type ('=' for the double bond, '#' for the triple bond).
Examples:
AB=BA - AB connected by a double bond with BA
AB#BA - AB connected by a triple bond with BA
2*AB - 2 repeats of AB, i.e., ABAB
2*S1 + 2*S2 - 2 repeats of S1 and 2 repeats of S2
Predicted molecular weight and length will be shown below.
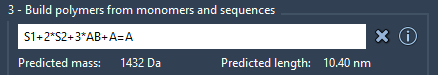
Click Generate polymer.
The app generates a polymer with a shape based on directions between the start and end atoms of each monomer. Excess hydrogens at connection sites are removed.

Advanced Options#
- Adjust Hydrogens on connection atoms – Automatically adjust (add/remove) hydrogens for each connection atom (start and end atoms of monomers) based on valence. This is also done to adjust their positions. If you would like to adjust hydrogens later, you can use Edit > Add hydrogens. You can also add hydrogens manually using the Add editor (see User Guide: Building molecules for more information).
- Adjust for clashes – Tries to avoid steric overlaps when connecting monomers (may cause slight curvature).
Tip
You can use a generated polymer as a new monomer to build block copolymers or extended chains.
Minimize the Polymer#
Use SAMSON's built-in minimizer: Edit > Minimize. Click again to stop minimization.
Note
SAMSON uses the Universal Force Field in its built-in minimizer.
Saving and Loading Projects#
If you would like to save/export the resulting polymer, select it and then click Home > File > Save selection as....
You can also save the full project to preserve monomer and sequence tables and load it later.
Use Cases#
- Build synthetic or biopolymers for material property prediction.
- Model conjugated polymers for electronic applications.
- Generate drug-polymer conjugates or molecular brushes.
- Use with simulation tools like GROMACS, LAMMPS, OpenMM, or SAMSON's own engines.
Related tutorials#
- Create molecular boxes with Molecular Box Builder for building larger molecular systems around generated molecules.
- GROMACS Wizard tutorials for preparing and simulating molecular systems.
- Generate crystal models for another materials and structure-building workflow.
Next step#
Save the generated polymer or continue preparing it for simulation with the toolchain that matches your system.
Need Help?#
Have questions or feedback? Feel free to reach out via the Forum, via e-mail, via the Feedback button in SAMSON, or by directly discussing with us.