Generate analogue series with positional analogue scanning#
Use positional analogue scanning in the SMILES Manager to generate analogue series directly in SAMSON from a starting molecule and a SMARTS pattern. This workflow is useful when you want to quickly explore substitutions, compare analogue properties, or prepare structures for docking and interaction analysis.
The feature was inspired by Pat Walters' blog post and the cited paper from Lewis Pennington, Ingo Muegge, and coworkers. In this tutorial, we use the same example molecule from that paper: CN1C=C(C(=O)Nc2ccc(-c3ccccc3)c(c2)C(F)(F)F)C(=O)c2ccccc12.
What you will learn#
In this tutorial, you will learn how to generate analogue series in SAMSON with positional analogue scanning in the SMILES Manager.
Before you start#
- Add the SMILES Manager extension.
- Open the extension and switch to the Positional Analogue Scanning tab before following the steps below.
- Use the companion SMILES Manager tutorial if you first need help importing or organizing SMILES data.
Start from a single molecule#
Let's begin by defining our starting molecule. You can do that in two different ways: either you enter the SMILES code of your molecule of interest or you select it in the SAMSON document and you press the Use selection button like in the gif below.
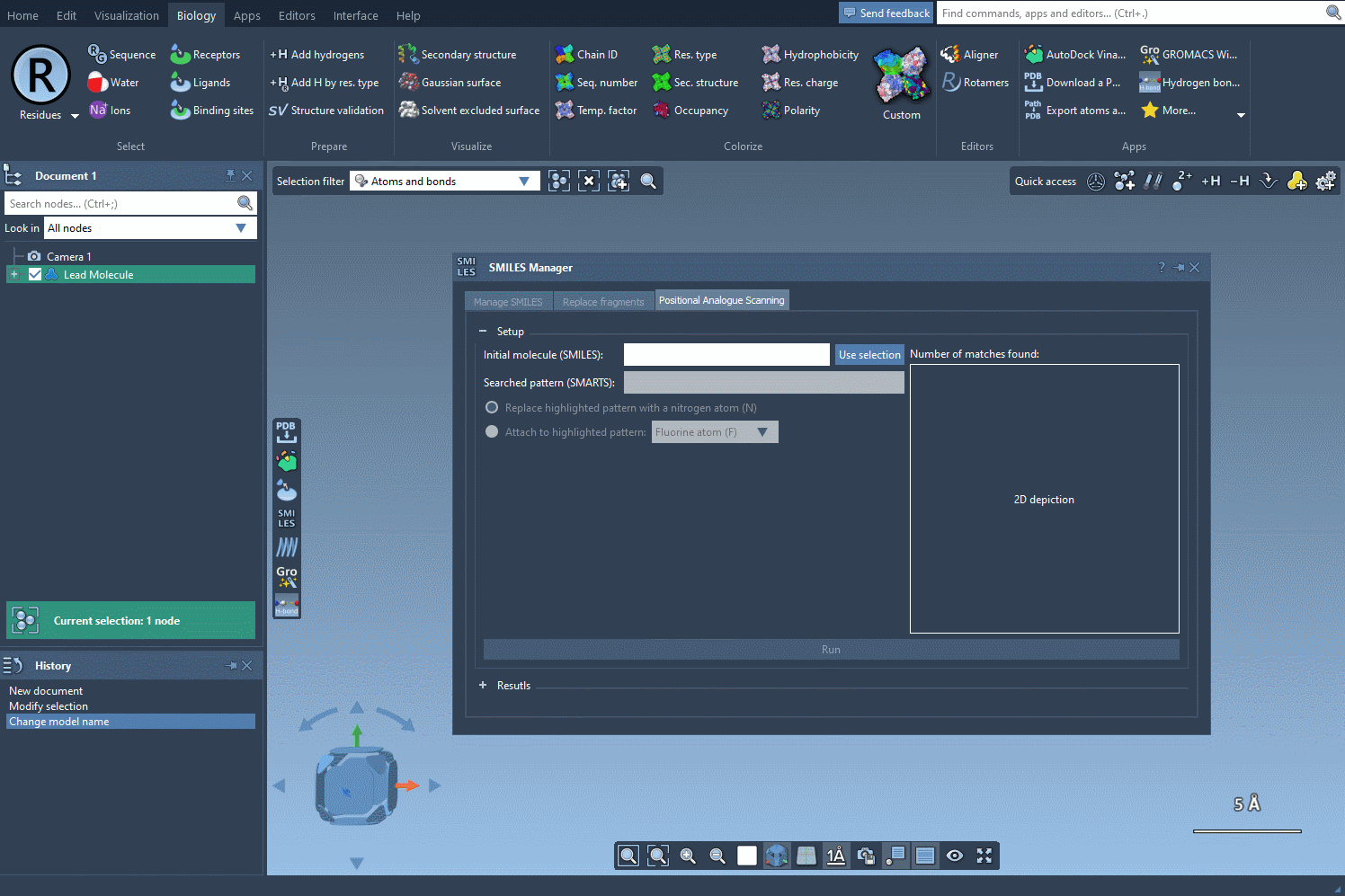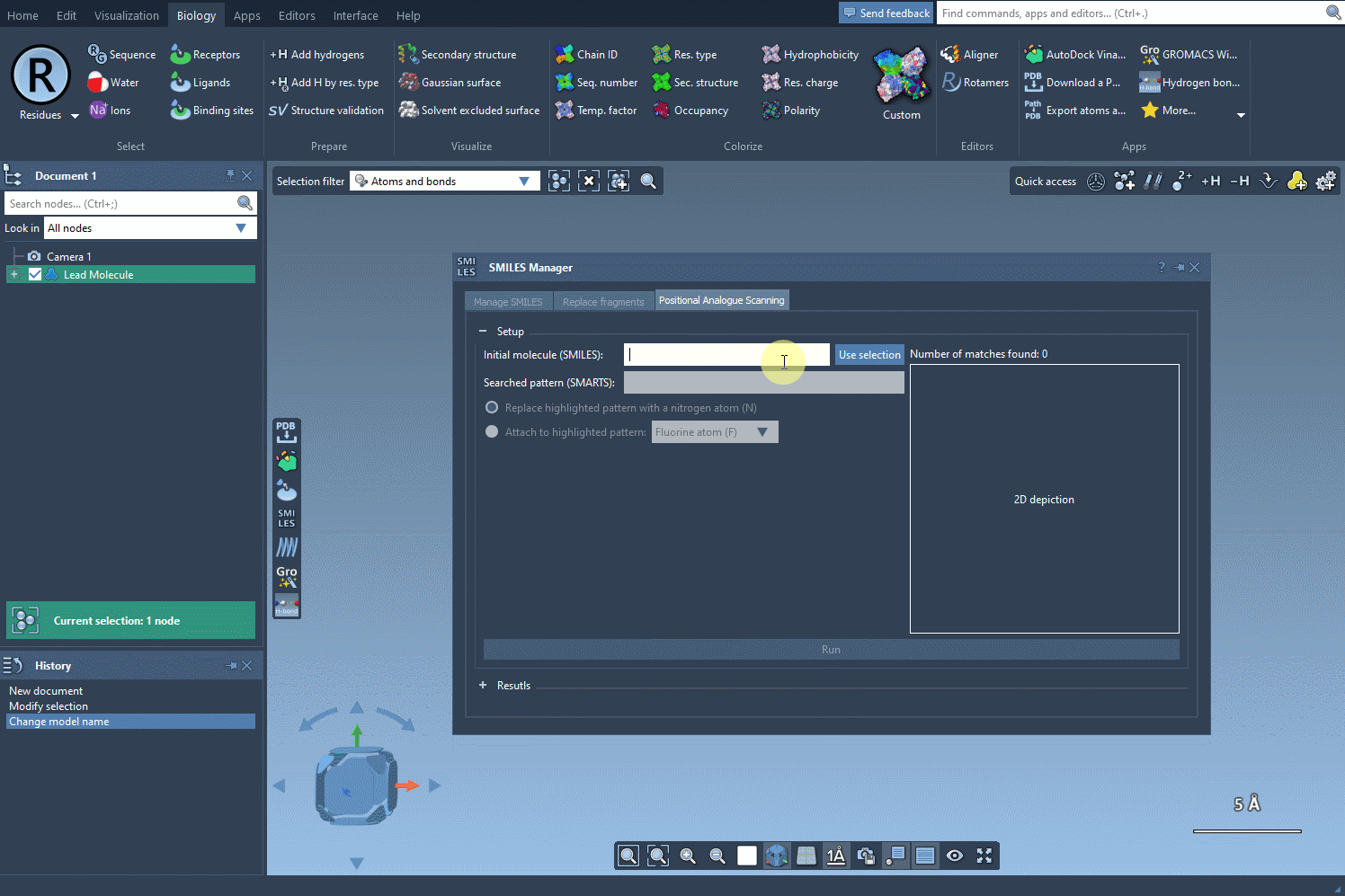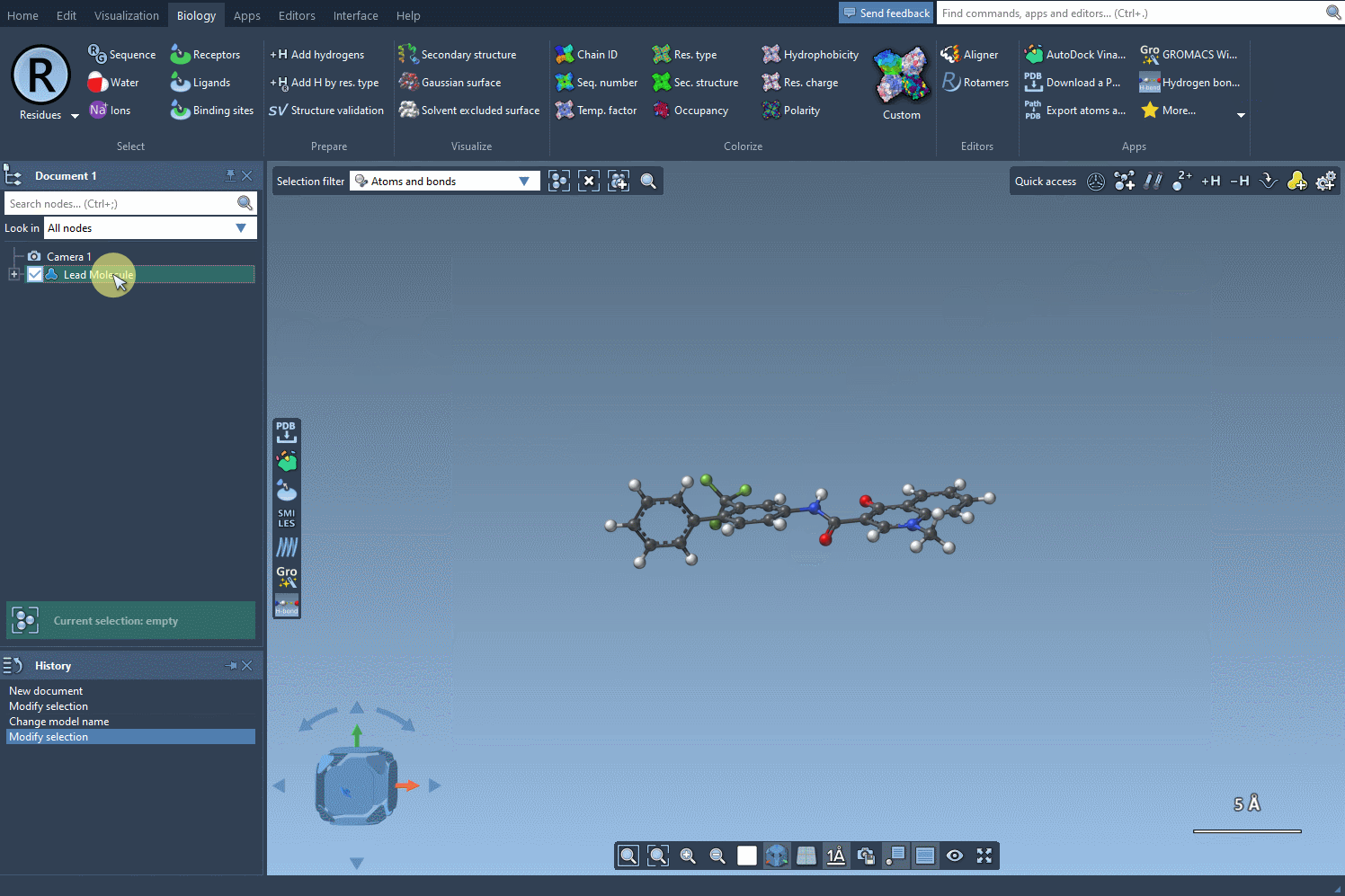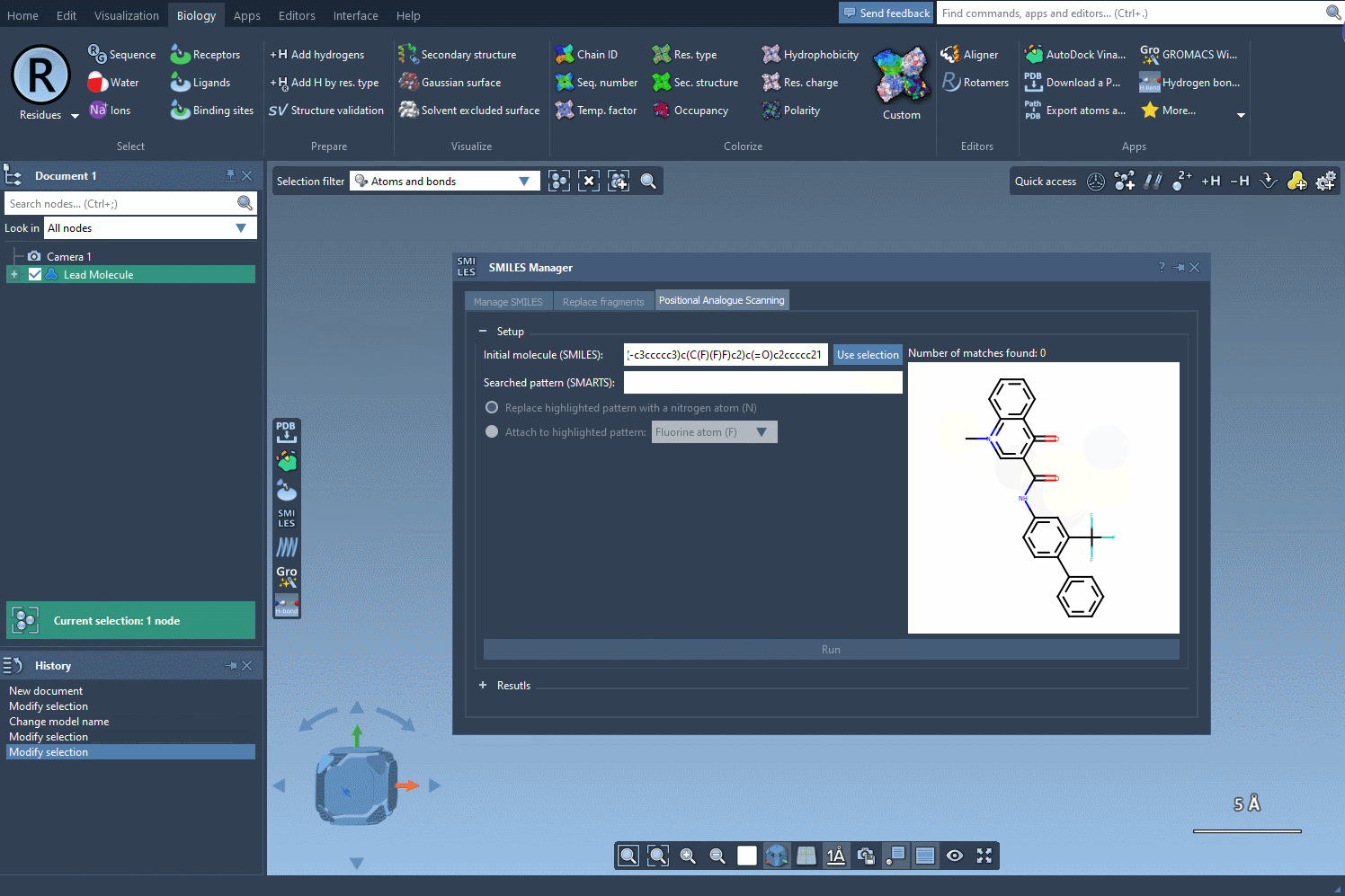
Define your searched pattern#
Search the initial structure for the pattern you want to replace or to which you want to attach a new atom or group. In our example, we will use aromatic carbon as a pattern by entering the SMARTS code [cH] in the corresponding field. As shown in the following gif, we can see that the found pattern in the initial molecule was highlighted a number of times.
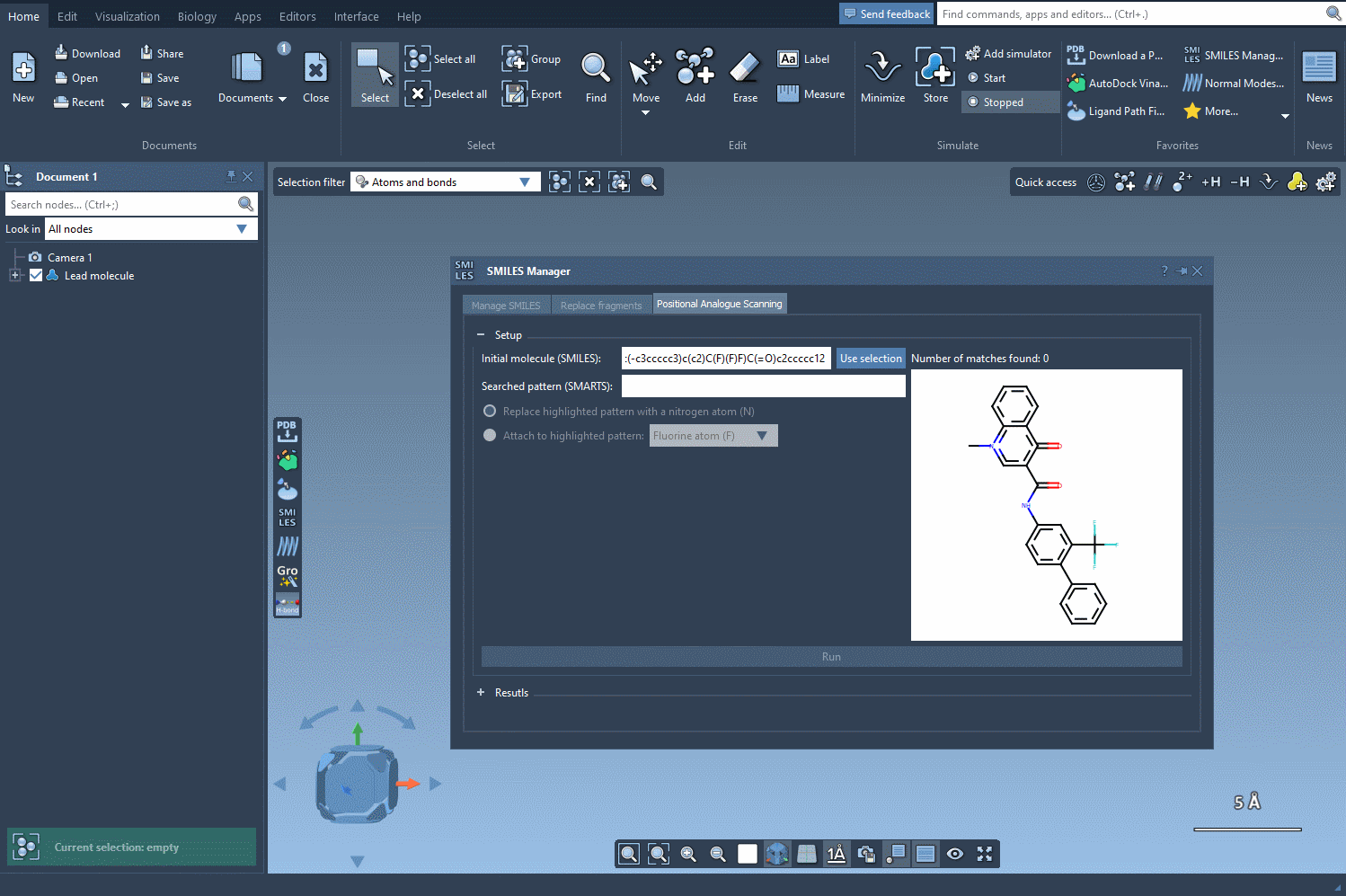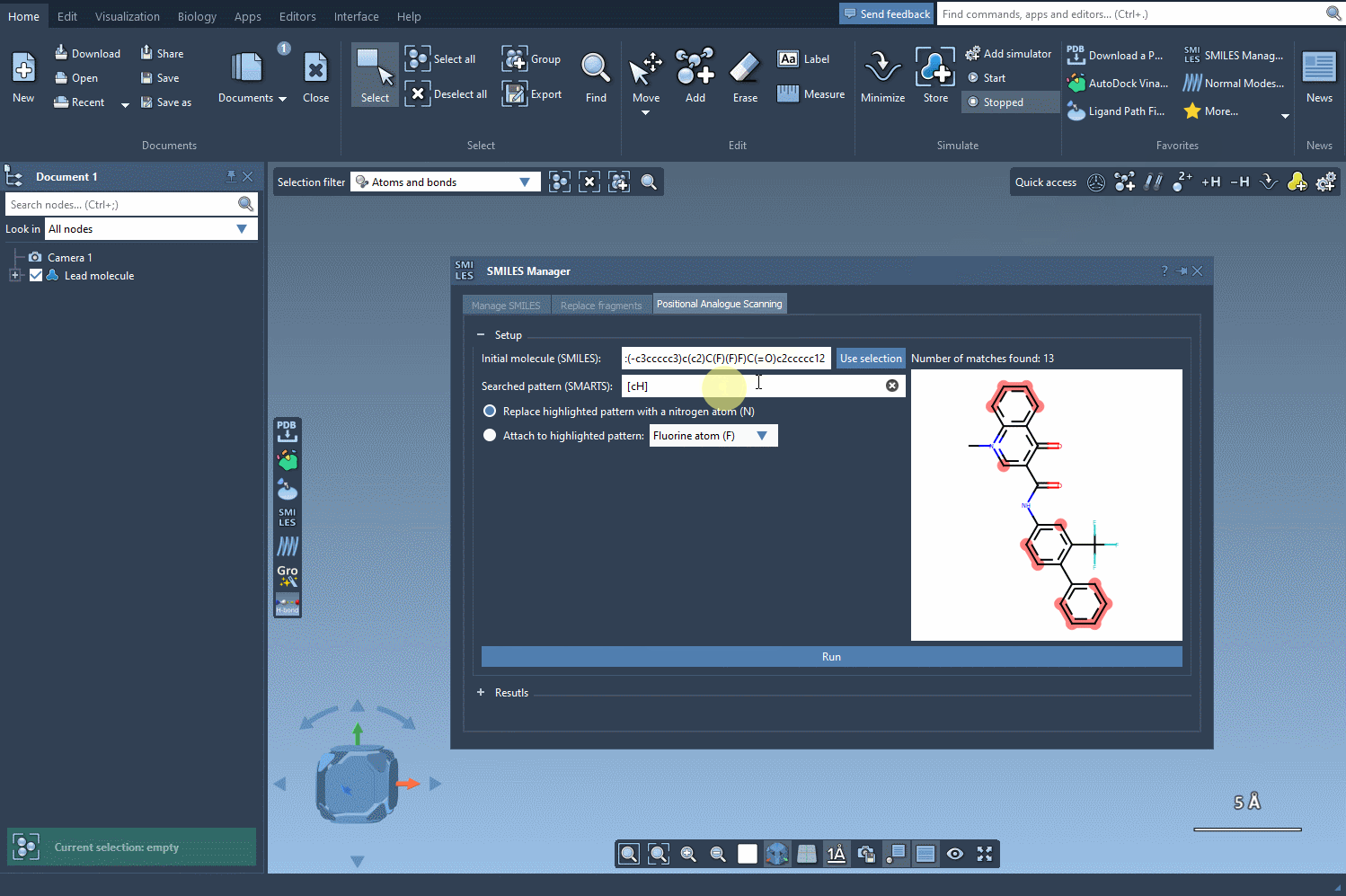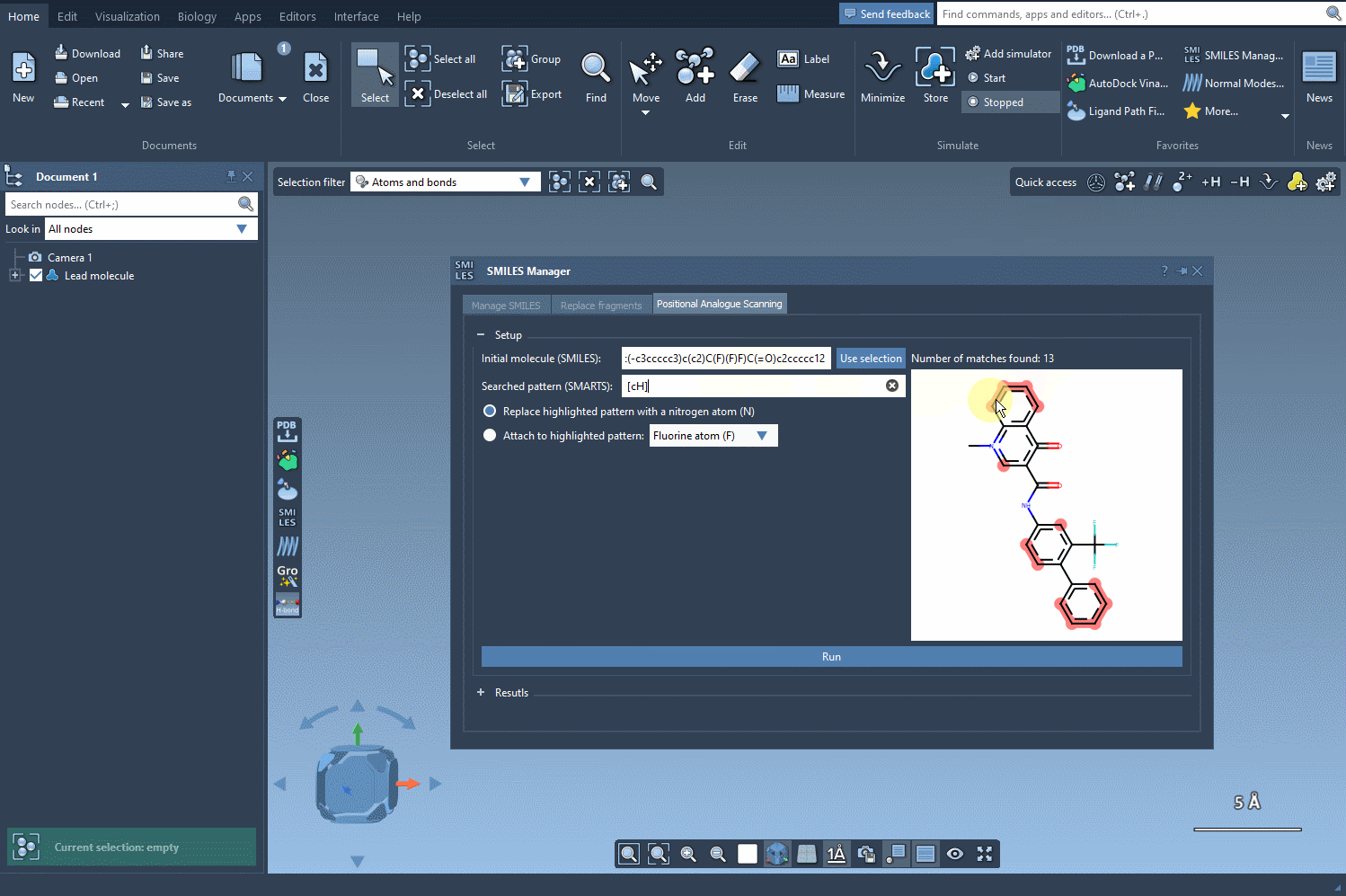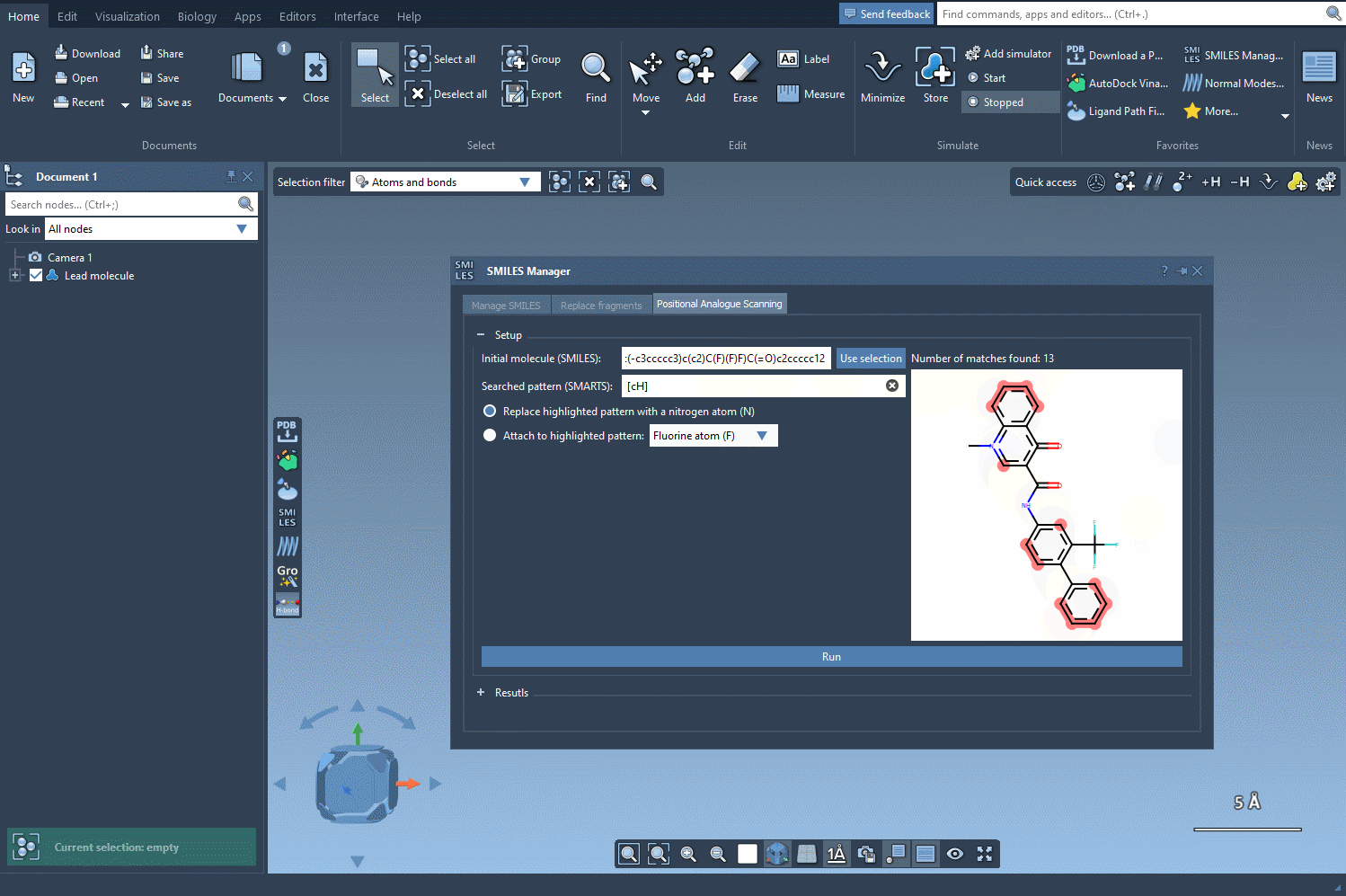
Replace or attach#
Now that we specified the pattern we want to scan, we choose the first option in order to replace it with a nitrogen atom (N). If you like, you can choose to attach to your pattern a fluorine atom (F) or a methyl group (CH3). Then we only have to click Run to generate the analogs and display them in the results section of the interface.
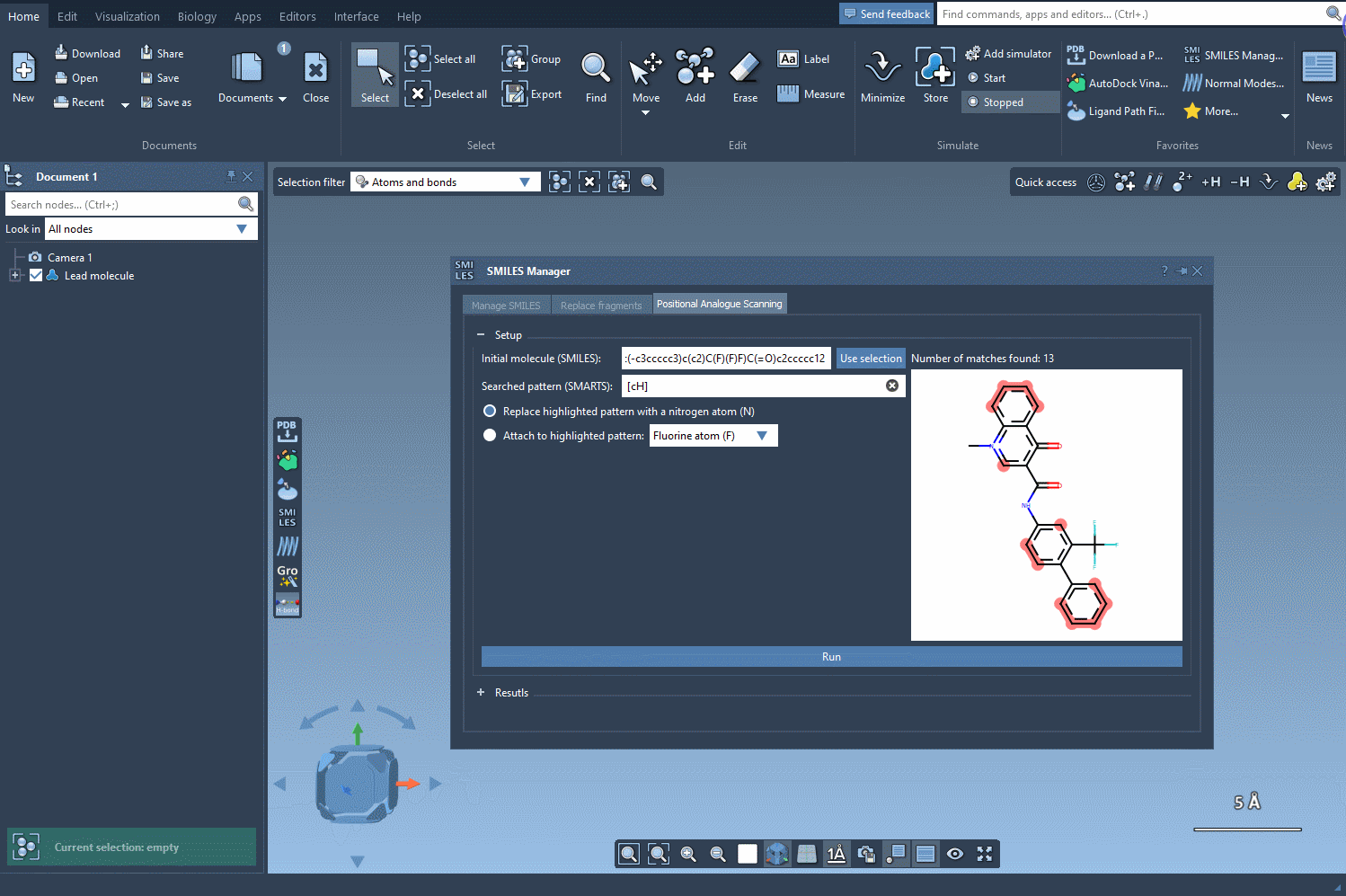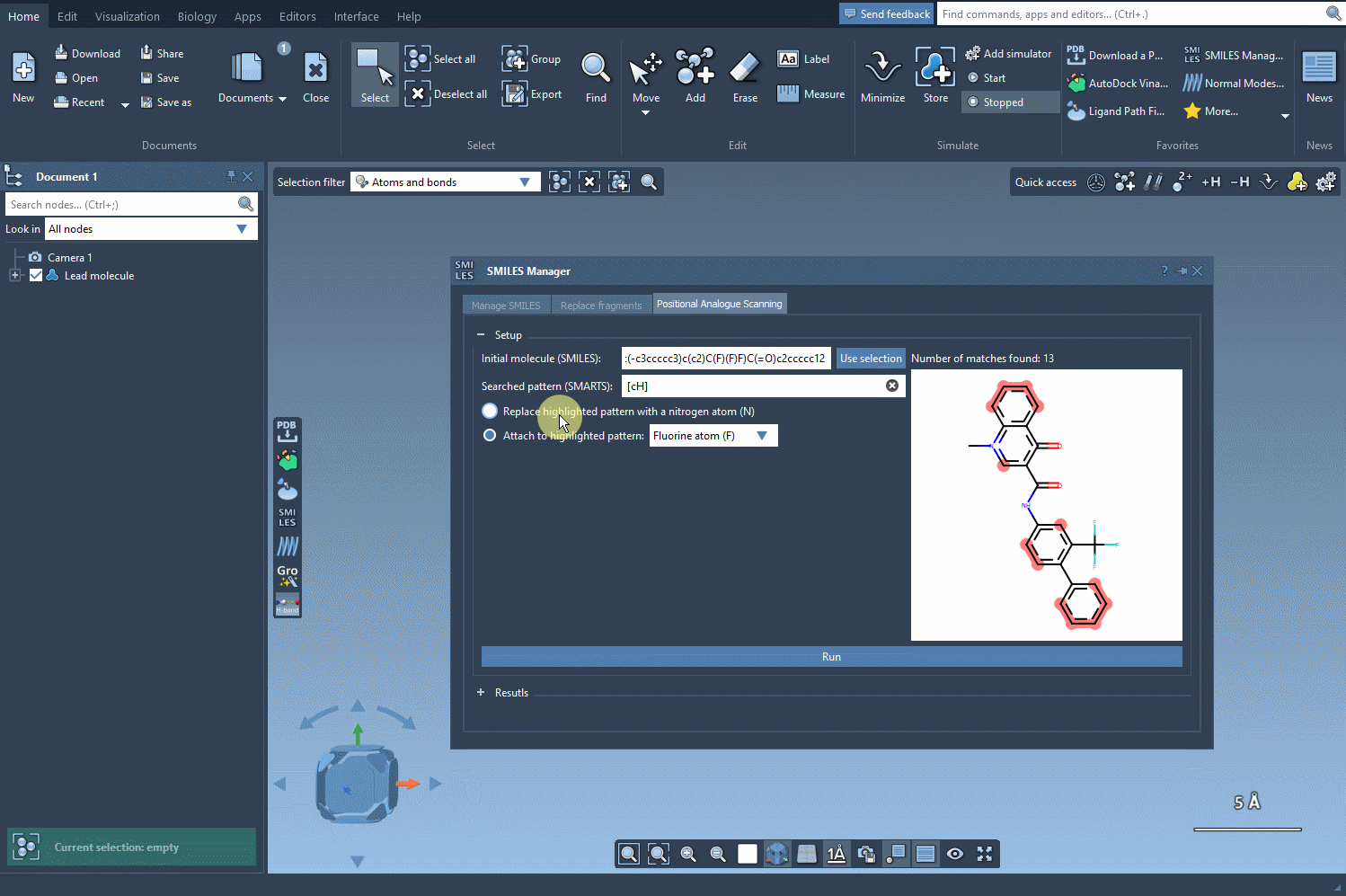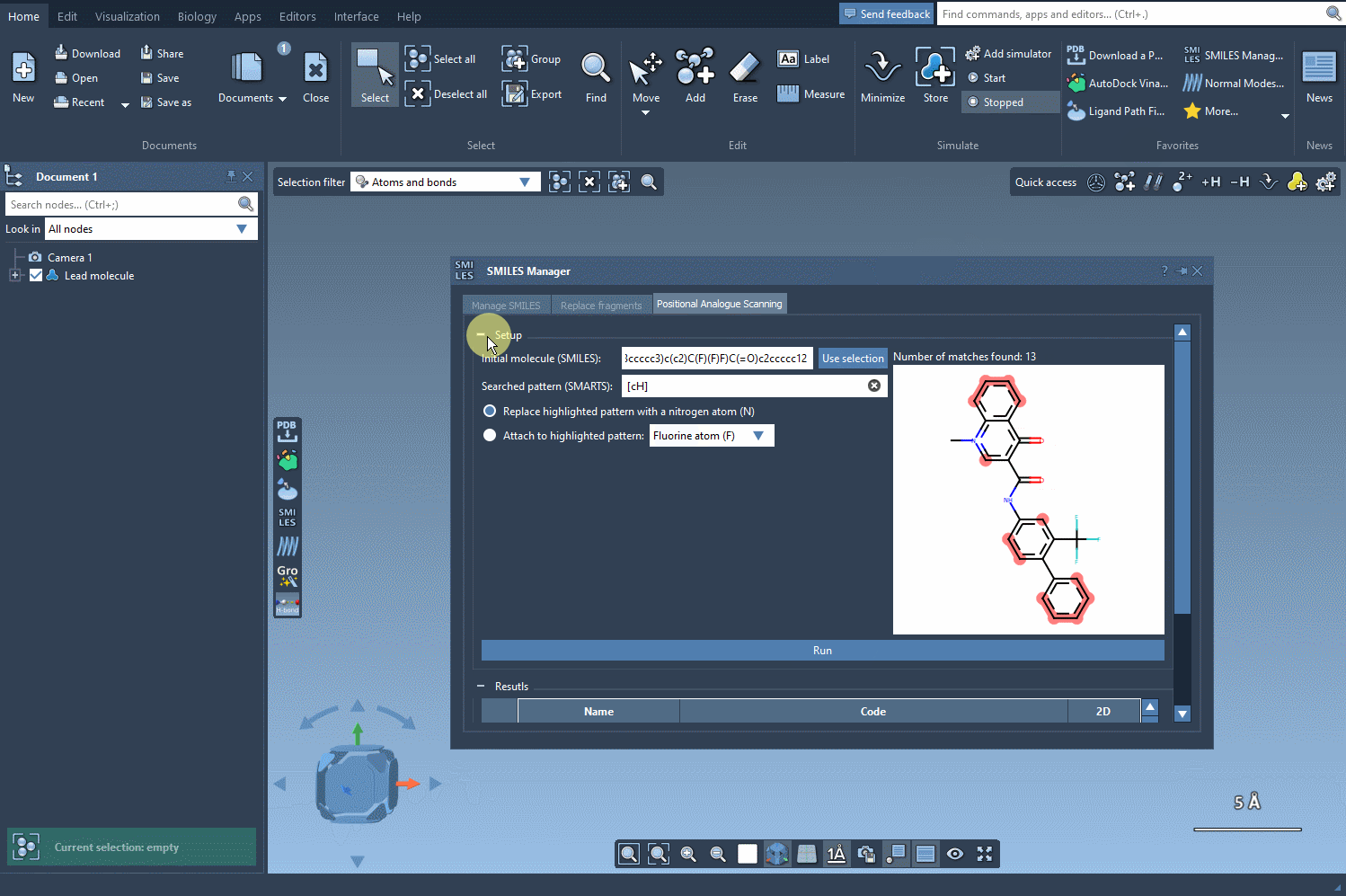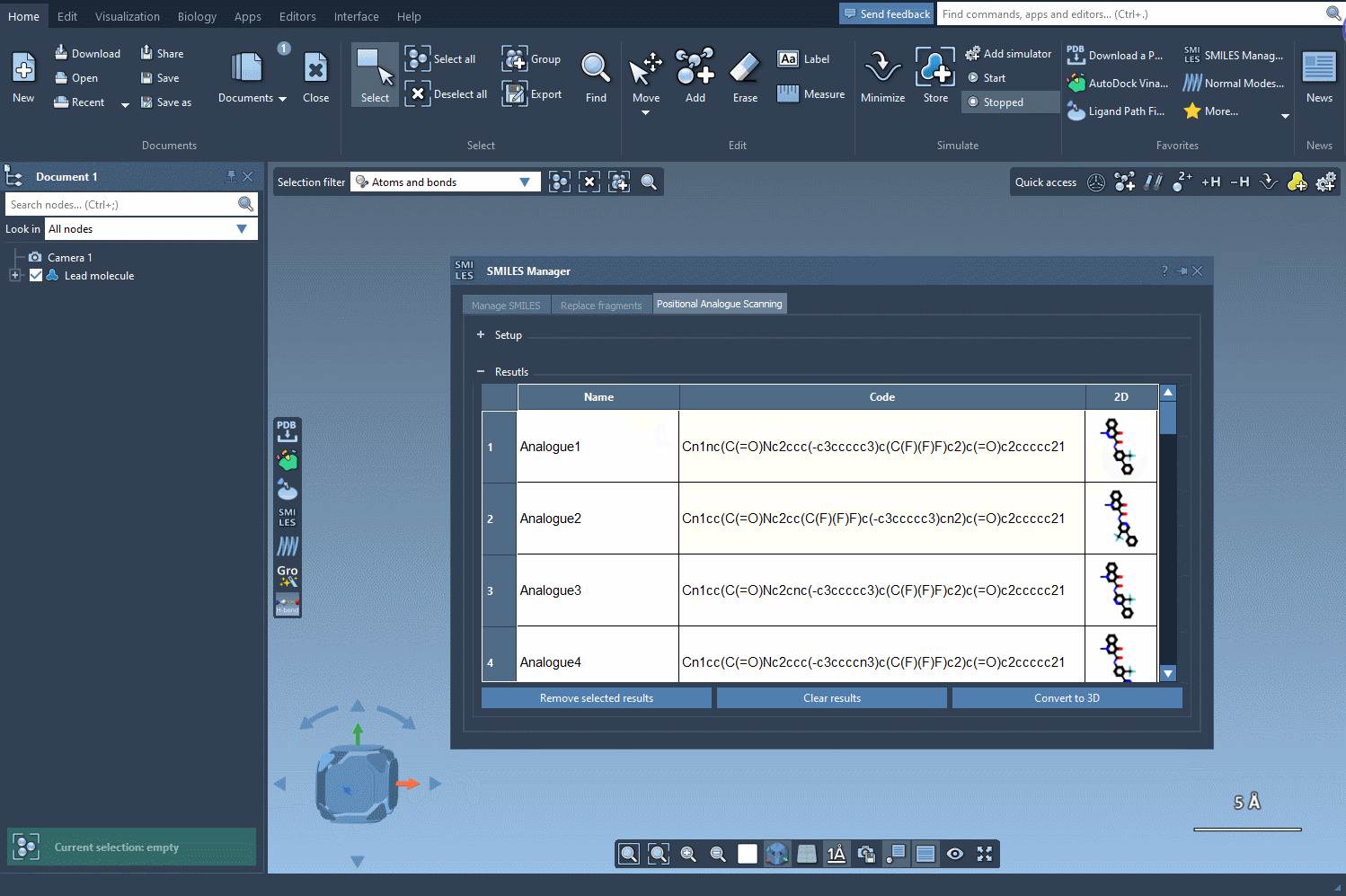
As you can see, it was quite fast to generate the SMILES codes and the 2D depiction of the analogs.
Modify your results#
In the results table you can do many actions:
- Modify the generated analogs: change the name or the SMILES code of the analogue of your choice by double-clicking on the corresponding cell in the results table.
- Open the 2D depiction image: double-click on the 2D depiction image to open it in a larger window. You can also do that by right-clicking on the image and clicking Open in the context menu.
- Hide/show the analogs' images in the results table: click on the 2D cell of the table's header to hide or show the 2D images in the results table.
- Generate a 3D structure of a specific analogue: right-click on the image of your analogue of interest and choose generate a 3D structure in the opened context menu.
- Remove only selected results: select the lines of the analogs you want to remove from the results table and click the Remove selected results button.
- Clear results table content: click on the Clear results button to remove all the generated analogs from the table.
Generate 3D structures of your analogs and study them#
Finally, let's generate the 3D structures of our generated analogs by clicking on the Convert to 3D button.
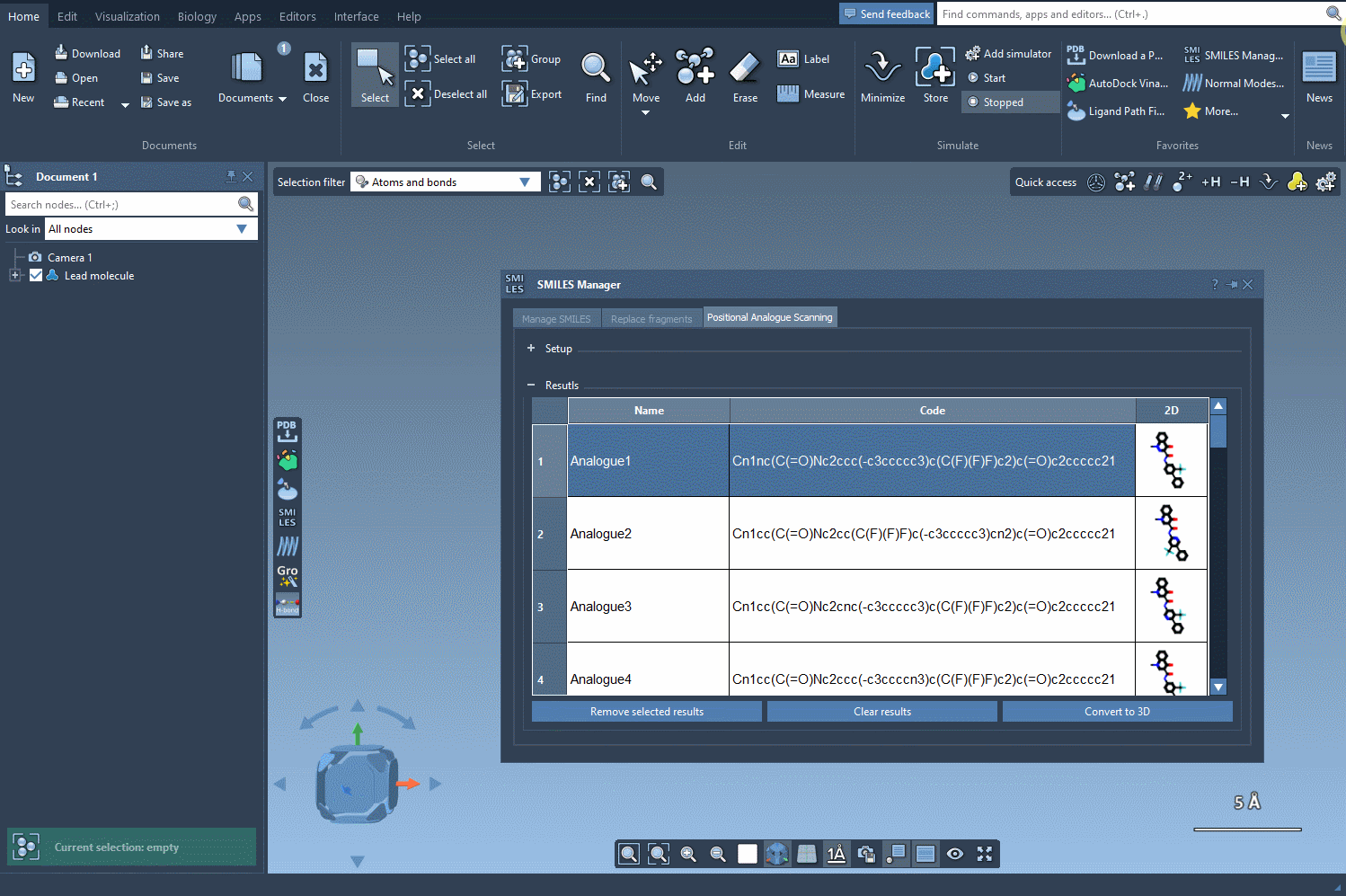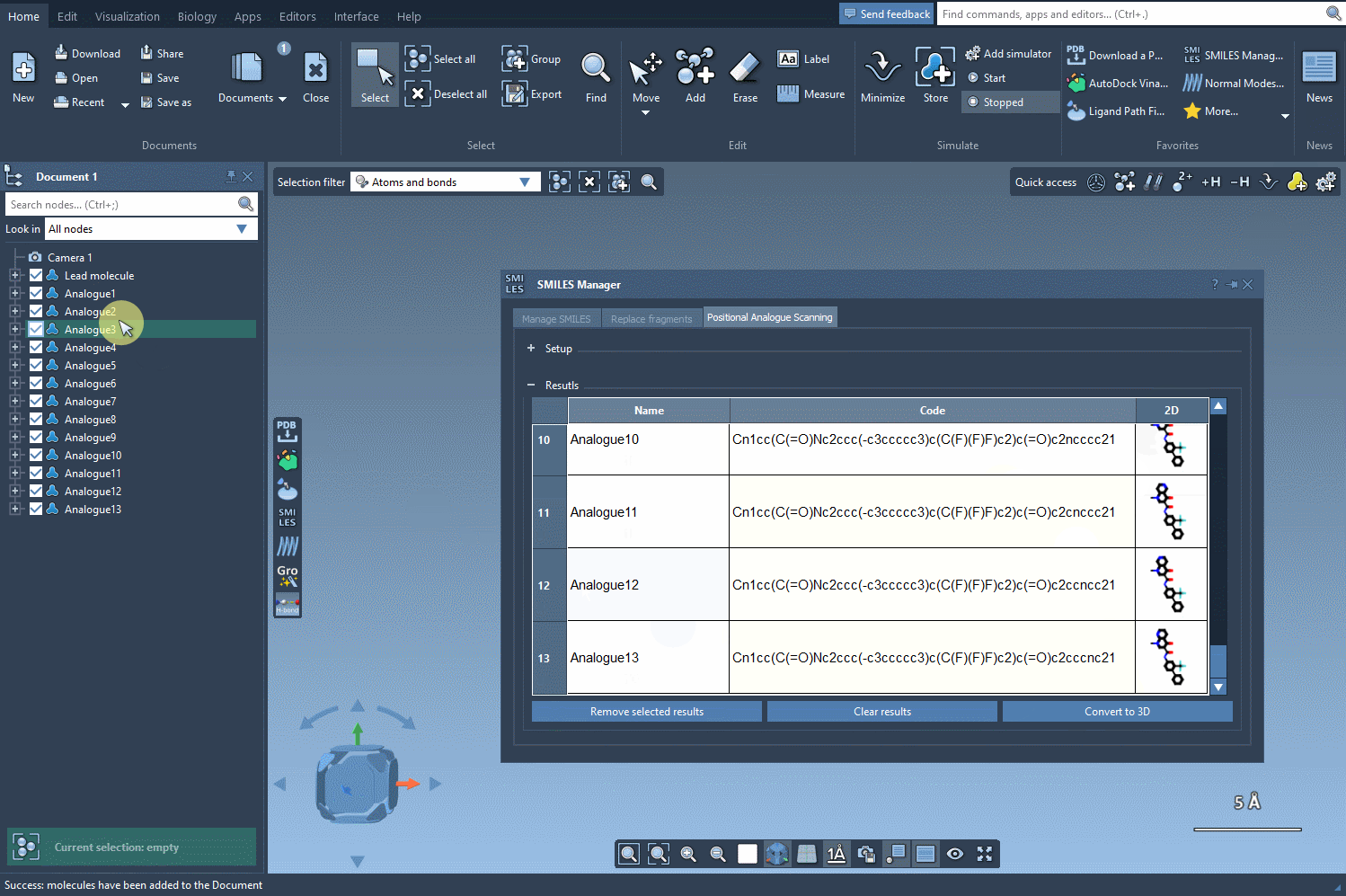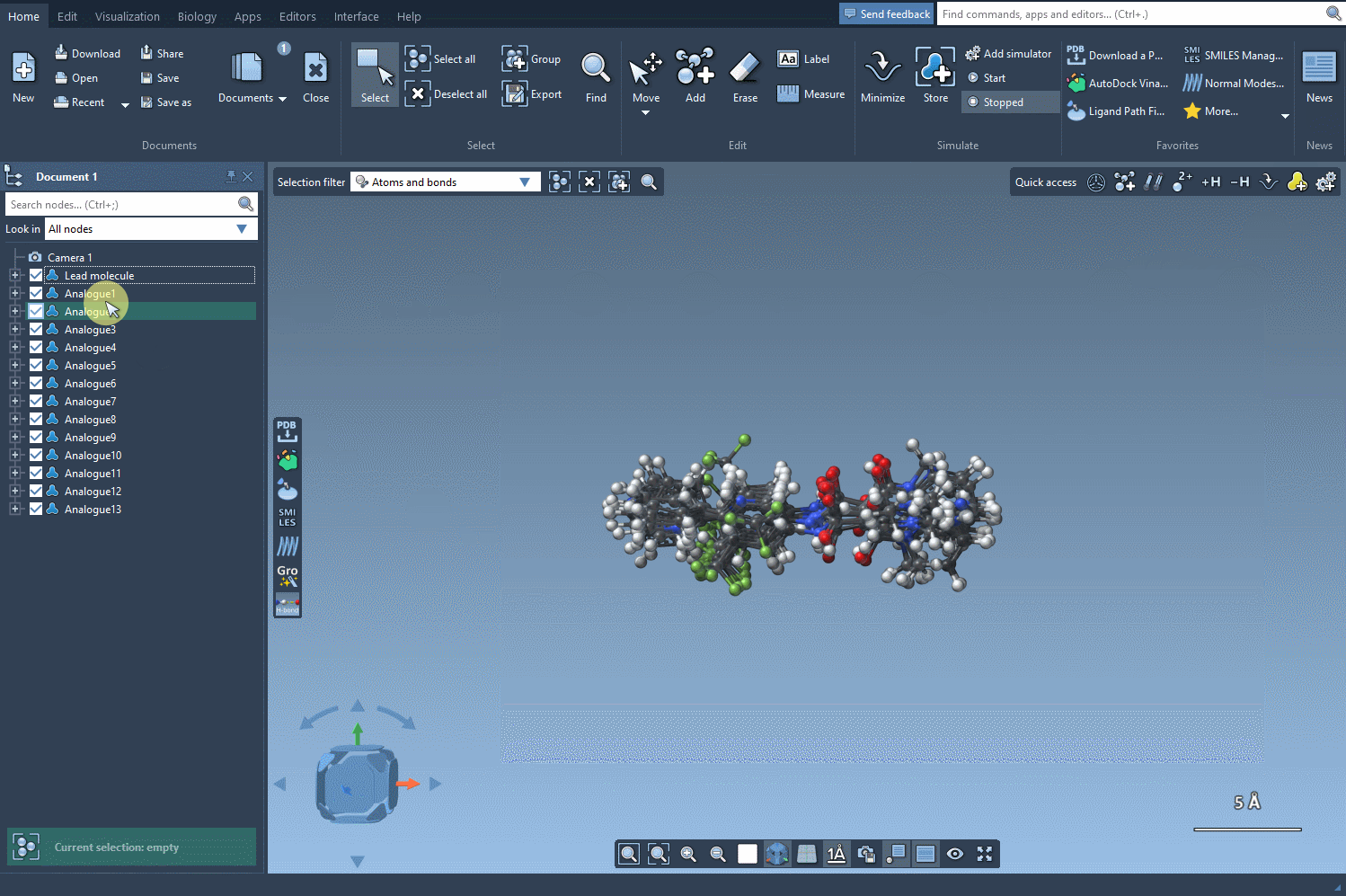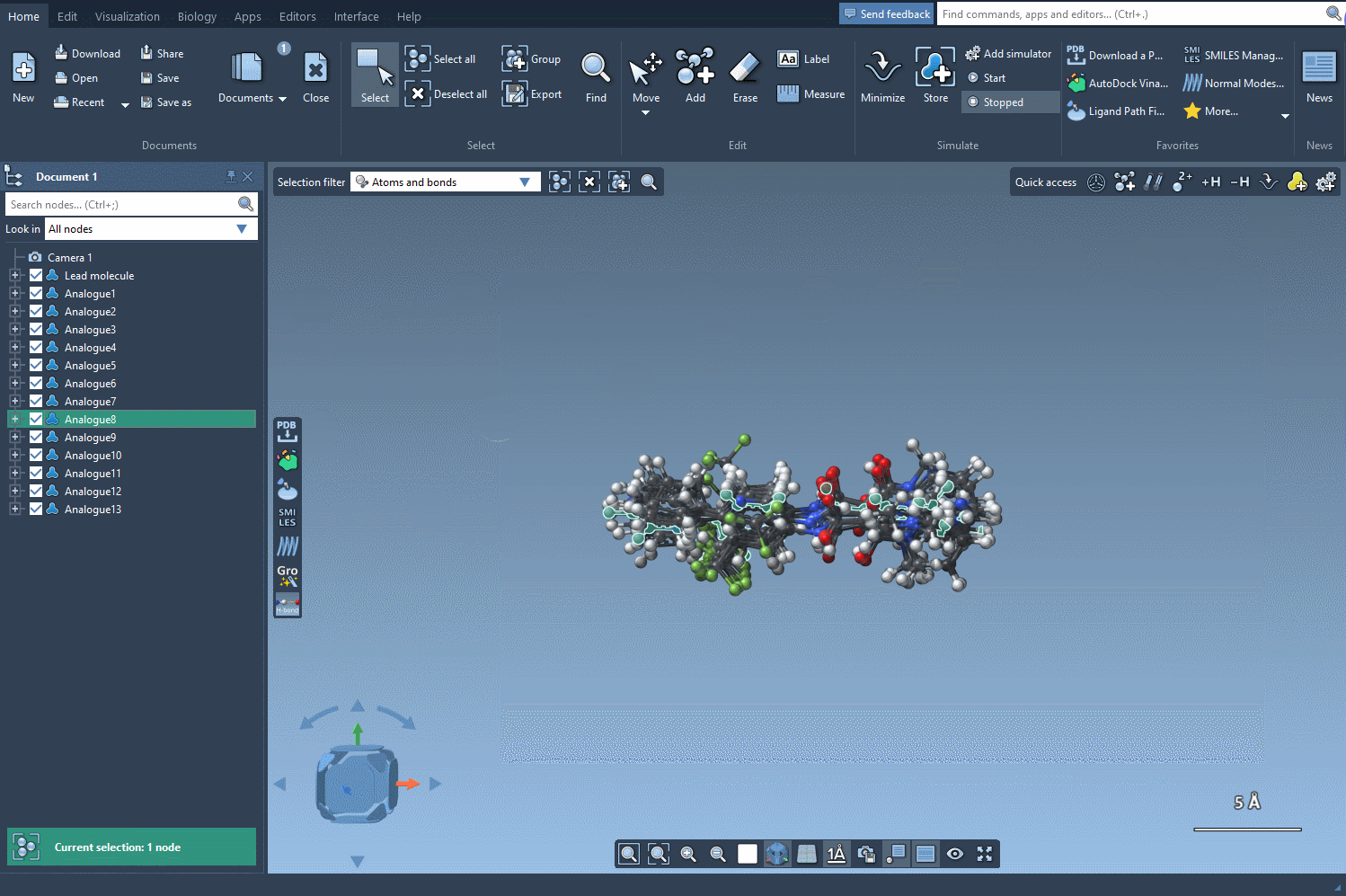
Now, you can proceed in the study of your molecules, for example, by performing a docking of the generated analogs using the Autodock Vina Extended extension. You can then examine which changes preserve key interactions with your target protein and which ones make new interactions.
Related tutorials#
- Using the RDKit - SMILES Manager for broader SMILES import, management, and fragment replacement workflows.
- Dock ligands and libraries with AutoDock Vina Extended for docking generated analogs.
- Strain Explorer for analyzing ligand strain in generated structures.
Next step#
Convert promising analogs to 3D structures, then dock, filter, or analyze them with the workflow that matches your design goal.
Need Help?#
Have questions or feedback? Feel free to reach out via the Forum, via e-mail, via the Feedback button in SAMSON, or by directly discussing with us.