Node Specification Language#
SAMSON has a powerful Node Specification Language (NSL) that can be used to select various structures (ligands, receptors, etc.) and nodes (atoms, residues, etc.) based on their properties. NSL can be used in the Find command or to filter nodes in the Document view.
Interactive tutorial in SAMSON (Help > Tutorials): "Selecting using the Node Specification Language".
What you will learn#
This page teaches the core NSL syntax, where to enter expressions in SAMSON, and which reference pages to use when you need more precise attribute queries.
NSL Quick Start#
This brief section introduces the fastest way to begin using the Node Specification Language (NSL) in SAMSON for search and filtering.
Use it when you are new to NSL and want a practical starting point before reading the full syntax and attribute-space references.
Where to use NSL#
- Select > Find for search-driven selections (see the Find command section to learn more)
- the filter box in the Document view for interactive filtering
First expressions to try#
node.type residueC or H"CA" within 5A of Snode.category ligand, receptor
Related pages#
Find command#
The Find command (1) in SAMSON lets you enter a NSL string to select nodes (atoms, residues, etc.) from the active document:
- Select > Find, , : Ctrl+F, : Cmd+F

When using the Find command, you may press the Tab key in the search box for context-sensitive completion. This is particularly useful when searching nodes by name. For example, entering "ALA (with the opening quote sign and without the closing quote sign to indicate a start of a name) and pressing the Tab key will list all nodes whose name begins with "ALA", e.g:
"ALA 22 Backbone""ALA 22 Side chain""ALA 22""ALA 28 Backbone""ALA 28 Side chain""ALA 28"...
Tip
You can also ask AI Assistant to generate an NSL expression for you - click on the ![]() Ask AI button on the left of the selection string. The AI Assistant knows the active document's hierarchy, so you can ask for selections specific to your document.
Ask AI button on the left of the selection string. The AI Assistant knows the active document's hierarchy, so you can ask for selections specific to your document.
Find using Document View#
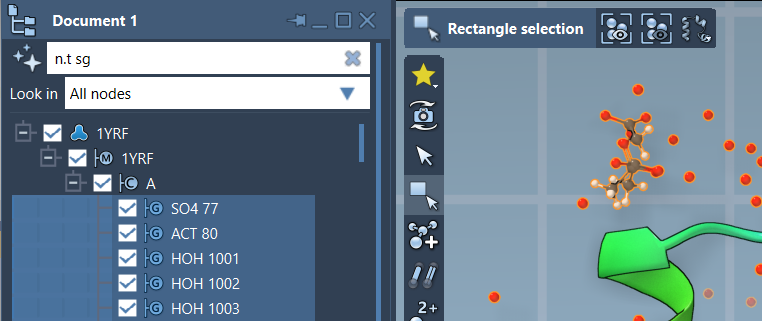
A NSL expression can also be used to filter nodes in the Document view (1) and to select them by clicking Enter, as shown in the image below when selecting structural groups via the following NSL: n.t sg (full version: node.type structuralGroup):
- Interface > Document view, , : Ctrl+1, : Cmd+1
Tip
You can also ask AI Assistant to generate an NSL expression for you - click on the ![]() Ask AI button on the left of the filter nodes in the Document view. The AI Assistant knows the active document's hierarchy, so you can ask for selections specific to your document.
Ask AI button on the left of the filter nodes in the Document view. The AI Assistant knows the active document's hierarchy, so you can ask for selections specific to your document.
Examples#
Here are some examples of NSL expressions:
node.category ligand, receptor(short version:n.c lig, rec): matches ligands and receptorsHydrogen(short version:H): matches hydrogensresidue.id 20:40, 50:60(short version:r.id 20:40, 50:60): matches residues with IDs between 20 and 40 and between 50 and 60atom.chainID > 2(short version:a.ci > 2): matches atoms with a chain ID strictly larger than 2Carbon in node.selected(short version:C in n.s): matches carbons in the current selectionbond.order > 1.5(short version:b.o > 1.5): matches bonds with order strictly larger than 1.5node.type backbone(short version:n.t bb): matches backbone nodes
Logical operators examples:
C or H: matches atoms that are carbons or hydrogensnode.type residue and not residue.type ALA(short version:n.t r and not r.t ALA): matches all non-alanine residues
Topology operators examples:
O in node.type sideChain(short version:O in n.t sc): matches oxygens in side chain nodesn.t a out of n.t r: matches atoms that do not belong to residuesnode.type sideChain having S(short version:n.t sc h S): matches side chain nodes that have at least one sulfur atomH linking O(short version:H l O): matches hydrogens bonded to oxygen atoms
Proximity operators examples:
"CA" within 5A of S(short version:"CA" w 5A of S): matches nodes named "CA" that are within 5 angstrom of any sulfur atom (use quotes, since names may contain spaces)node.type residue beyond 5A of node.selected(short version:n.t r b 5A of n.s): matches residue nodes beyond 5 angstrom of the current selectionresidue.secondaryStructure helix(short version:r.ss h): matches residue nodes in alpha helices
Tip
Specification by attributes#
Nodes and groups of nodes may be specified by their attributes.
SAMSON's NSL supports different attribute spaces based on node types. Since nodes may have different types in SAMSON (e.g. atom, bond, etc.), each attribute is defined in an attribute space:
| Attribute space | Short name |
|---|---|
node |
n |
animation |
an |
atom |
a |
backbone |
bb |
bond |
b |
camera |
ca |
chain |
c |
conformation |
co |
file |
fi |
folder |
f |
label |
la |
light |
li |
mesh |
me |
molecule |
mol |
nodeGroup |
ng |
note |
nt |
path |
p |
presentation |
pr |
propertyModel |
pm |
renderPreset |
rp |
residue |
r |
segment |
s |
sideChain |
sc |
structuralGroup |
sg |
structuralModel |
sm |
visualModel |
vm |
The node attribute space (short name: n) corresponds to attributes that are defined for each node. For example, the selection flag is a node attribute, since each node has a selection flag. Hence, the NSL expression node.selectionFlag true may match any node whose selection flag is true, regardless of its node type (atom, bond, etc.). For convenience, the other attribute spaces inherit relevant attributes from the node attribute space. So, you can also use atom.selectionFlag true to match all atoms whose selection flag is true.
Specification by name#
Nodes may be specified by their names using strings enclosed with quotes.
A quoted string can be used in three ways:
- exact matching: a quoted string without wildcards matches the full name exactly
- glob matching: use
*and?inside a quoted string for simple pattern matching - regular-expression matching: prefix a quoted string with
reto use a full regular expression
Note
Bond names are formed from the names of the atoms they bond, and cannot be searched by name.
Exact matching#
A quoted string without wildcards matches the full node name exactly.
Examples:
"ALA": matches nodes named exactlyALA"ALA 22": matches nodes named exactlyALA 22"CA": matches nodes named exactlyCA
Glob matching#
For simple name patterns, quoted strings support the following wildcards:
*: matches zero or more characters?: matches exactly one character
This is often the easiest way to search by name when you only need simple patterns.
For example, entering "ALA*" may yield ALA residues, backbones, and side chains, e.g:
"ALA 22 Backbone""ALA 22 Side chain""ALA 22""ALA 28 Backbone""ALA 28 Side chain""ALA 28"...
Examples:
"CA*": matches nodes whose names start withCA"*Backbone": matches nodes whose names end withBackbone"*AL*": matches nodes that containALin their names, e.g. ALA and VAL residues, backbones, and side chains"C?": matches names such asCAorCBorCG, etc."???": matches names made of exactly three characters"ALA ?2": matches names where exactly one character appears before2, for exampleALA 12
Tip
Use regular expressions for more complex cases.
Regular expressions#
For more advanced matching, prefix the string with re, for example re"..." or re "...".
These expressions use the Perl-like regular-expression syntax (supported by Qt's QRegularExpression). Regular expressions are useful when * and ? are not enough, for example when you want to match one of several names, restrict the number of characters, or specify character ranges.
Examples:
re"ALA.*"orre "ALA.*": matches names that begin withALAre".*Backbone$": matches names that end withBackbonere"^(CB|CG|CE)$": matches exactlyCB,CG, orCDre"^[A-Z]{3}$": matches names made of exactly three uppercase letters, which is useful for many residue-name patternsre"^Chain [A-C]$": matches names such asChain A,Chain B, orChain Cre"^[A-Za-z]+ [0-9]+$": matches names made of a word followed by a number
Tip
Use glob matching for simple cases, and use re only when you need more advanced matching.
Specification by symbols and element names#
In order to efficiently match atoms of a given element type, symbols and element names are valid NSL expressions.
Possible values: atomic symbols and element names (with the first character capitalized).
Examples:
Carbon: matches carbon atomsC: matches carbon atomsCa: matches calcium atomsH: matches hydrogen atoms
Lists and ranges#
NSL supports providing lists (divided by comma ,) and ranges (via colon :) wherever possible. Examples:
atom.chain A, B, C(short version:a.c A,B,C): matches atoms in chains A, B, and Cresidue.id 20:40, 50:60(short version:r.id 20:40, 50:60): matches residues with IDs between 20 and 40, and between 50 and 60atom.x -1nm:1nm(short version:a.x -1nm:1nm): matches atoms with x-coordinate between -1 nm and 1 nm
Operations on sets#
Logical operators#
Logical operators may be used to perform operations on sets.
Available operators:
andnotorxor(exclusive or).
Examples:
sg.id 1000:1040 and sg.nat < 4: matches structural groups with IDs between 1000 and 1040 that have less than 4 atomsa.sn <= 20 or a.sn >=40: matches atoms with serial number smaller than 20 or larger than 40n.t r and not r.t CYS: matches residues that are not cysteinsa.sn >= 20 xor a.oc >= 0.5: matches atoms which either have a serial number larger than 20, or have an occupancy larger than 0.5, but not those that satisfy both conditions
Note
Without proper care, the not condition might produce surprising results. For example, not r.t CYS does not return the list of residues that are not cysteins. Indeed, a folder is also a node which is not a residue that's a cystein. If only residues should be returned, a proper query would be n.t r and not r.t CYS.
Topology operators#
Containment operators may be used to specify inclusion in or out of a set, one set having nodes from another set, or having atoms linked to atoms in another set.
Available operators:
inout ofhaving(short name:h) - whether a node has another node(s)linking(short name:l) - whether an atom or a structural node is linked to another atom(s) or structural node(s) containing atoms
Examples:
n.t a in "2AZ8": matches atoms in "2AZ8"n.t a in r.t CYS: matches atoms that belong to cysteinsH in r.t ARG: matches hydrogens that belong to argininsH in n.h: matches hidden hydrogensn.t a out of r.t PRO: matches atoms that do not belong to prolinesnode.type sideChain having S(short version:n.t sc h S): matches side chain nodes that have at least one sulfur atomH linking O(short version:H l O): matches hydrogens bonded to oxygen atomsnode.type residue linking (residue.type PRO, CYS)(short version:n.t r l (r.t PRO, CYS)): matches residues that have atoms linked to atoms in PRO or CYS residues, this includes PRO and CYS residues themselves.
Proximity operators#
Proximity operators may be used to select nodes based on distances.
| Operator | Short version |
|---|---|
within {distance} of |
w {distance} of |
beyond {distance} of |
b {distance} of |
Examples:
C within 5A of "GLN 2"(short version:C w 5A of "GLN 2"): matches carbons within 5 angstrom of "GLN 2"node.type atom beyond 5A of "2AZ8-IA"(short version:n.t a b 5A of "2AZ8-IA"): matches atoms beyond 5 angstrom of "2AZ8-IA"
Note
Without proper care, the within and beyond conditions might produce surprising results. For example, * within 5A of "GLN 2" will select the document node, since some atoms in the document are within 5 angstrom of "GLN 2": the atoms that belong to "GLN 2". If only atom nodes should be returned, a proper query would be n.t a within 5A of "GLN 2". Note that this latter query would also return atoms that belong to "GLN 2". If only atoms outside "GLN 2" should be returned, then a proper query would be n.t a within 5A of "GLN 2" out of "GLN 2".
Topological proximity operators#
Topological proximity operators may be used to select nodes based on the maximum number of bonds between atoms.
| Operator | Short version |
|---|---|
withinBond {number_of_bonds} of |
wb {number_of_bonds} of |
Examples:
atom.symbol C withinBond 3 of atom.symbol S(short version:a.s C wb 3 of a.s S): matches Carbon atoms that are within 3 or less bonds of Sulfur atomsresidue.type ALA withinBond 4 of residue.type GLY(short version:r.t ALA wb 4 of r.t GLY): matches alanine residues that are within 4 or less bonds of glycine residues
Length units#
| Length unit | Short name |
|---|---|
femtometer |
fm |
picometer |
pm |
angstrom |
A |
nanometer |
nm |
micrometer |
um |
millimeter |
mm |
centimeter |
cm |
meter |
m |
Mass units#
| Mass unit | Short name |
|---|---|
dalton |
Da |
kilodalton |
kDa |
megadalton |
MDa |
gigadalton |
GDa |
electronMass |
auMass |
yoctogram |
yg |
zeptogram |
zg |
attogram |
ag |
femtogram |
fg |
picogram |
pg |
nanogram |
ng |
microgram |
ug |
gram |
g |
kilogram |
kg |